Produced in Partnership with RedShiftBioReviewed by Maria OsipovaFeb 14 2024
Matrix Metalloproteinases (MMPs) are currently an important area of research. Exploring these proteins not only enhances our comprehension of protein biology but also sparks significant interest in creating specific MMP inhibitors for therapeutic purposes.
To support these research objectives, information about MMP structure is crucial. This study used Microfluidic Modulation Spectroscopy (MMS) to evaluate the structure of MMP-2, MMP-8, and MMP-9 using RedShift BioAnalytics’ first and second-generation instruments to compare the quality of the data obtained.
The structural similarities and differences between each of these proteins were also investigated. Although two out of the three MMPs share similar functions, the findings revealed distinct structures for all three.
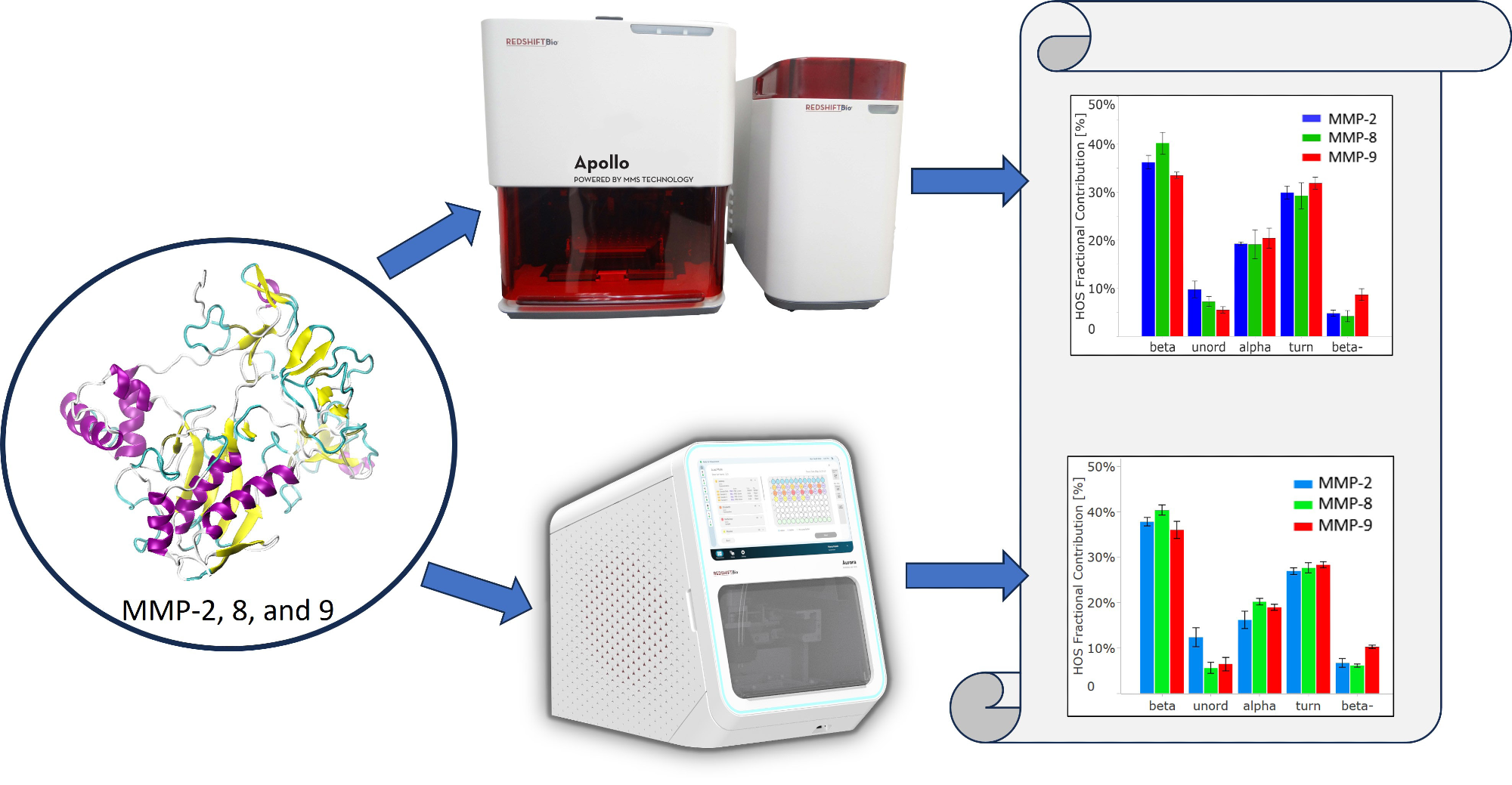

Image Credit: RedShiftBio
Introduction
MMPs are a class of proteases responsible for degrading extracellular matrix (ECM) proteins. Overexpression of specific MMPs is associated with various pathological disorders, including multiple sclerosis, cancer, and strokes.1
More recently, elevated MMPs have been observed in COVID-19 patients due to their important roles in lung physiology.2,3 As a result, there is substantial research interest in using MMPs as potential biomarkers for predicting COVID-19 severity and creating inhibitors for targeted treatment in severe cases.4
Comprehending the structural variations among different MMPs is critical for enhancing this research objective. Various MMPs play specific roles in ECM tissue remodeling. For instance, MMP-8 (also called collagenase 2) digests collagens, while MMP-2 and MMP-9 (known as gelatinases A and B, respectively) target gelatins.
All these MMPs exhibit augmented expression levels in COVID-19 patients.2,3 Therefore, assessing both the commonalities and differences in the structure of these MMPs will prove instrumental in this context.
MMP-2, -8, and -9 have been extensively investigated and structurally characterized, especially in their catalytic forms. The crystalline structures of the catalytic domains of these proteins are depicted in Figure 1.6-8 The structures of all three proteins are very similar.
The catalytic domains comprise three alpha-helices and five strands of beta-sheets in their secondary structure. Figure 2 displays the full-length proteins, or proenzymes, of MMP-2 and -9.
Both the pro-domains and the catalytic domains share significant similarities in their structures. However, the fibronectin domains show significant differences and are primarily comprised of random coils (unordered structures). Currently, the crystalline structure of the full-length MMP-8 remains unresolved.
This study utilized microfluidic MMS to characterize and compare the structures of the proenzymes of these MMPs.
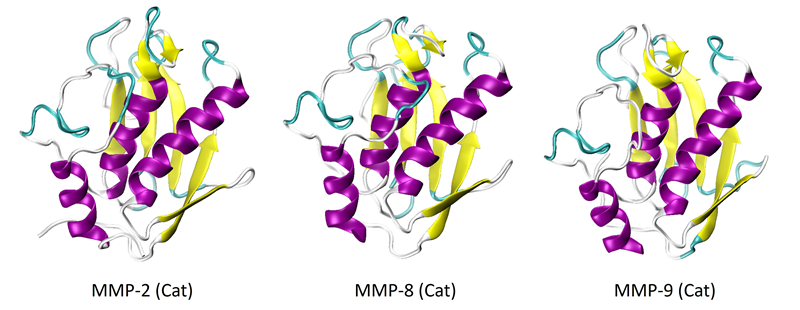
Figure 1. X-Ray crystal structures of the catalytic domains of MMP-2 (PDB: 1QIB), MMP-8 (PDB: 2OY4), and MMP-9 (PDB: 1GKC). Image Credit: RedShiftBio
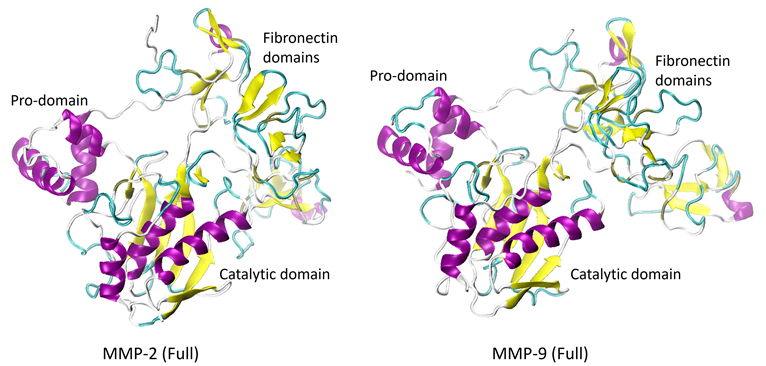
Figure 2. X-Ray crystal structures of the full-length proenzymes of MMP-2 (PDB: 1CK7) and MMP-9 (PDB: 1L6J). The C-terminal hemopexin-like domain is not shown. Image Credit: RedShiftBio
MMS probes the amide I band of the infrared (IR) spectrum to discern protein structure. To guarantee precise and real-time background subtraction, MMS constantly modulates against the reference buffer. The sensitivity offered by this method makes it particularly valuable for quality control and is well-suited for use with various formulation buffers.
This study used the first-generation MMS system, Apollo, along with RedShift BioAnalytics’ second-generation instrument, Aurora, which substantially reduces the volume required.
Both instruments are equipped with a high-power Quantum Cascade Laser, which, in comparison to conventional Fourier transform IR (FTIR) light sources, is considerably more intense.
This enhanced light intensity, in combination with modulating background subtraction, renders MMS approximately thirty times more sensitive than FTIR and five times more sensitive than circular dichroism (CD) for the detection of small changes in protein structure.5
Methods
One milligram of each MMP-2, MMP-8, and MMP-9 was acquired from SinoBiological (Wayne, PA).
To ensure buffer consistency, MMP-2 and MMP-9 underwent dialysis against their respective formulation buffers of 50 mM tris, 150 mM NaCl, 10 mM CaCl2, and 0.05 % Brij35 at pH 7.5, while MMP-8 was dialyzed against phosphate buffer saline (PBS).
Each sample, diluted to 1 mg/mL with a volume of 1 mL, was analyzed in triplicate using a first-generation Apollo MMS system. For comparison, the same samples were also analyzed on the second-generation Aurora system.
The Aurora instrument necessitates only 50 µL of sample. The Apollo system requires approximately 700 µL of sample.
In both systems, a backing pressure of 5 psi was applied to transfer the samples into the flow cell. Following this, samples were modulated at 1 Hz between the sample and reference buffer (using the same buffer utilized during dialysis) for background subtraction.
The differential absorbance was quantified within the range of 1588- 1711 cm-1. Replicates were averaged, and all samples were normalized to attain the absolute absorbance spectra.
Data processing in this study adhered to the procedures outlined in RedShift BioAnalytics’ previous application notes. In summary, raw differential absorbance data was transformed into absolute absorbance, normalized by concentration and pathlength. The second derivatives of the absolute absorbance spectra were then computed to increase spectral features.
The generated plot was inverted and baselined to create a similarity plot, qualifying the area of overlap when compared to a control, offering a measure of similarity between samples.
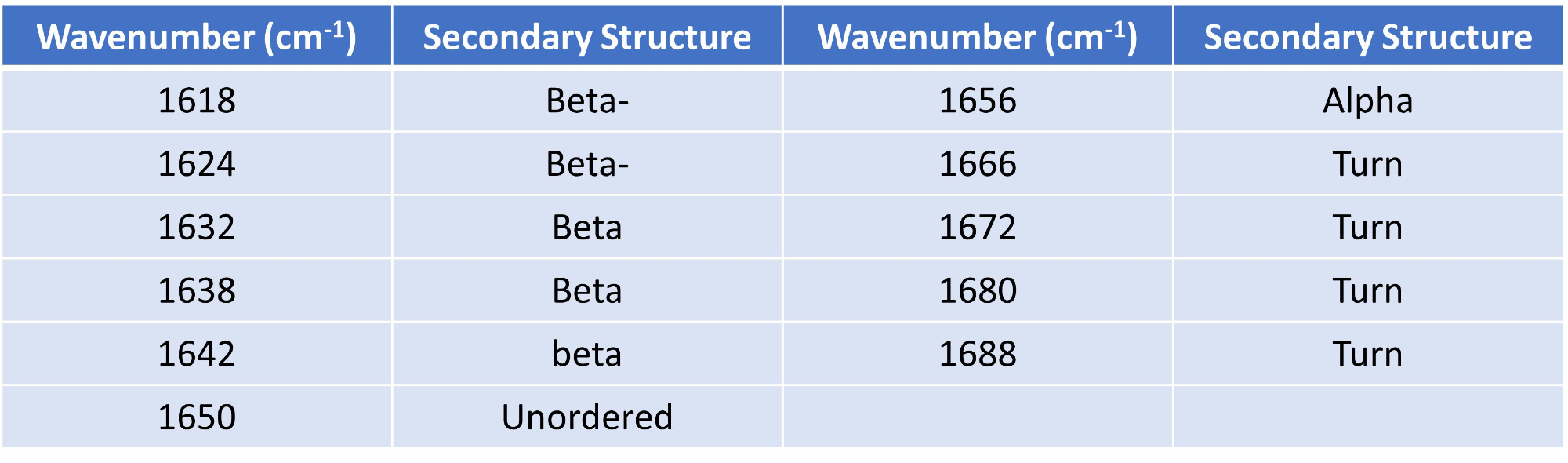
Using Gaussian curve fitting, 11 Gaussians were fitted, and the higher order structure (HOS) was calculated based on the determination of different secondary structural elements across the amide I band (shown in Table 1).
Table 1. Gaussian curve fit settings and HOS structural element designations. Source: RedShiftBio
Results and Discussion
The MMS findings highlight substantial structural differences among the three MMP subtypes, MMP-2, MMP-8, and MMP-9 (Figure 3).
Despite all three proteins exhibiting a mixture of alpha-helix (peaks at 1653-1658 cm-1) and beta-sheet (peaks at 1636-1640 cm-1) structures, as depicted in Figure 3A, their peak positions and intensities differ noticeably.
All three spectra exhibit less intense peaks at 1683-1687 cm-1 and 1618-1624 cm-1, representing the beta-turn and intermolecular beta-sheet structures, respectively.
Contrary to the expectations of MMP-2 and MMP-9 having more similar structures because of their functional likeness, the MMS findings suggest otherwise. MMP-2 and MMP-9 show the most substantial shifts, around 5 wavenumbers, for both the alpha-helix and beta-sheet peaks, with MMP-8 falling between the other two.
Regarding intermolecular beta-sheet peaks, MMP-9 shows a considerably more intense peak at 1624 cm-1, 6 wavenumbers higher than MMP-2 and MMP-8 at around 1618 cm-1.
The relative abundance of these secondary structural elements in each protein was determined using a Gaussian curve fitting with the delta software and is shown in the HOS bar chart in Figure 3B.
In all three proteins, the most prevalent secondary structure is beta-sheet, followed by beta-turn and alpha-helix.
In line with the similarity spectra, MMP-8 contains the most beta-sheet structures among the three proteins. MMP-2 exhibits the most unordered structures, and MMP-9 displays the most intermolecular beta-sheet structures (indicated as “beta-”).
Although they have similar biological functions, MMP-2 and MMP-9 do not share as much structural likeness as compared to MMP-8.
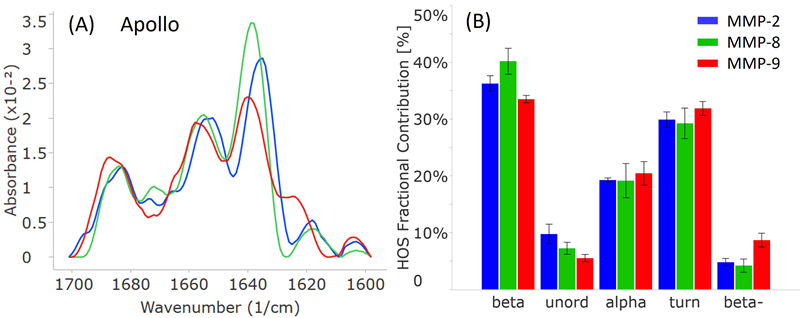
Figure 3. Apollo data showcasing the secondary structural comparison of MMP-2, MMP-8, and MMP-9. (A) Similarity plots (baselined second derivative spectra) illustrating the peaks within the amide-I band for each MMP. (B) HOS plot displaying the relative abundance of each secondary structural element for the MMPs. The error bars represent +/- the standard deviation (N=3). Image Credit: RedShiftBio
For comparison, all three MMP samples underwent analysis on RedShift BioAnalytics’ second-generation instrument, Aurora. As depicted in Figure 4A, the distinct peak locations and intensities align with the Apollo data.
While the Aurora spectra may seem less smooth, particularly around 1680 cm-1, this is attributed to the increased resolution from 4 cm-1 (Apollo) to 1 cm-1 (Aurora), resulting in higher resolution and less interpolation in the Aurora spectra.
The HOS bar chart in Figure 4B corroborates the breakdown of HOS and attests to the quality of data for Aurora. The error bars and repeatability, as shown in Table 2, demonstrate that Aurora provides slightly higher data quality.
This is because of RedShift BioAnalytics’ ability to have higher signal averaging with lower volume consumed per replicate.
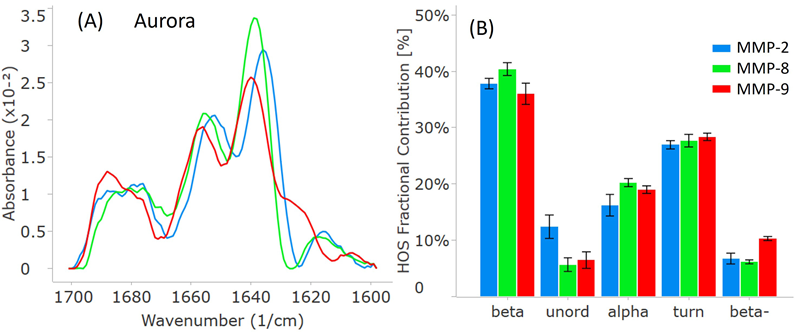
Figure 4. MMP secondary structure plots from the second-generation Aurora instrument. (A) Similarity plots (baselined second derivative spectra) illustrating the peaks within the amide-I band for each MMP. (B) HOS plot displaying the relative abundance of each secondary structural element for the MMPs. The error bars represent +/- the standard deviation (N=3). Image Credit: RedShiftBio
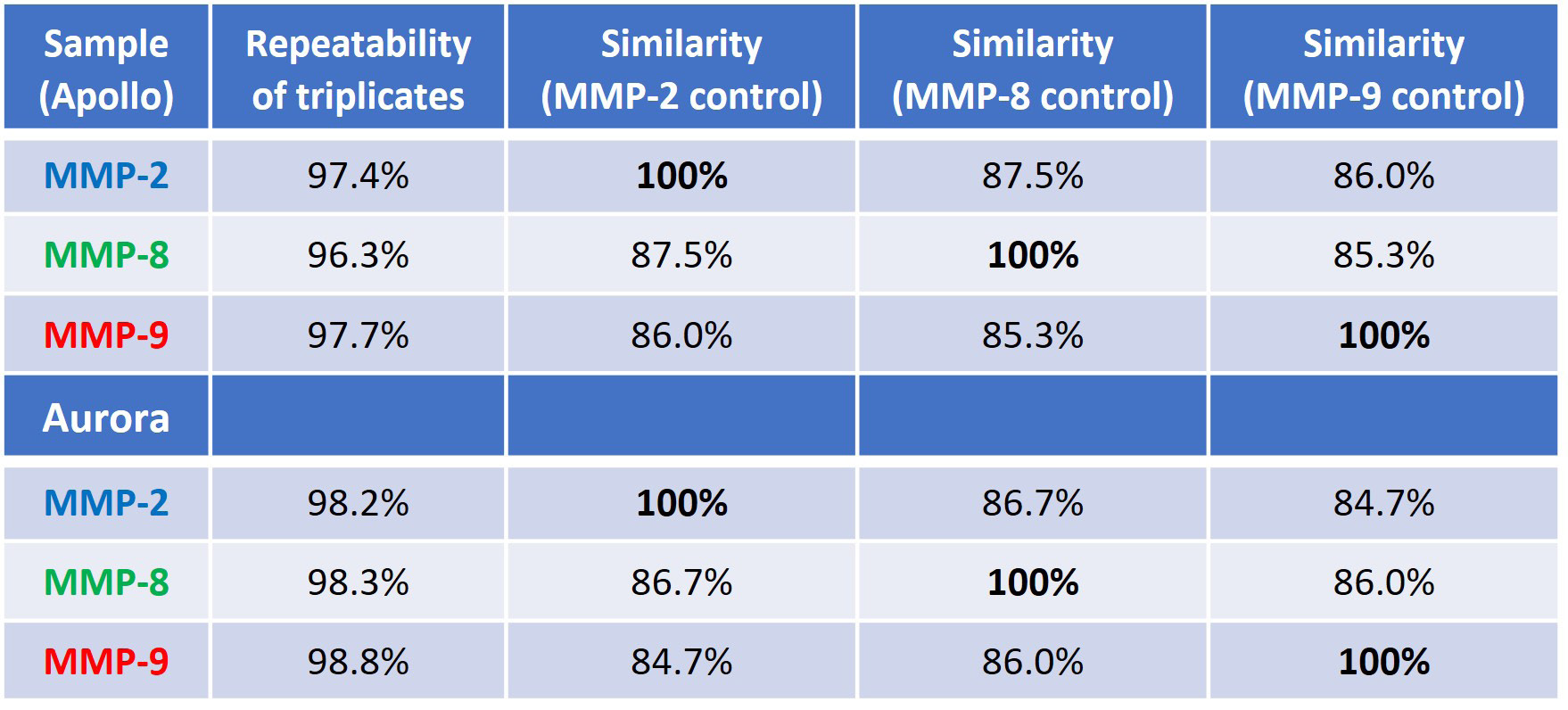
The measurement of structural similarity among the MMP proteins involves quantifying the area of overlap (AO) between the similarity plots shown in Figures 3A and 4A. Table 2 offers information on the repeatability of the triplicate measurements within each sample and the sample-to-sample likeness in terms of percentage AO.
For a detailed comparison, each MMP protein served as a control sample (indicated by 100 % similarity) with the first similarity column using MMP-2, the second column using MMP-8, and the third column using MMP-9 as the reference.
This underscores the concept of proteins being "similarly different" from one another. In other words, two proteins can share the same percent similarity to a third protein, but they can still possess distinct structures themselves, as evident in this study.
Table 2 shows that, regardless of the protein used as a reference, the other two proteins consistently exhibit relatively low similarity scores (84-87 % sample-to-sample similarity with 96-97 % replicate repeatability, and >98 % for Aurora), indicating once again that all three proteins have different structures.
Table 2. Repeatability of measurement and sample-to-sample similarity (the control for each similarity comparison is set at 100%). The top portion of the table is data from Apollo, and the bottom is comparing data from Aurora. Source: RedShiftBio
Most MMP isoforms, including those examined in this study, possess a C-terminal hemopexin-like domain in their full-length protein following expression (proenzyme). As depicted in Figure 5, showing MMP-9 as a dimer, this domain exhibits a four-bladed beta-propeller structure comprised of symmetrical blade-shaped beta sheets.
These beta-propeller structures are typically known for their involvement in protein-protein interactions.
Consequently, MMP-9 often exists in dimeric form. In this data, the detection of a small amount of intermolecular beta-sheet signals at 1618 and 1624 cm-1 indicates the possibility of dimerization or, more broadly, protein-protein interactions in all the samples in solution.
This underscores MMS’ capability to study protein-protein interactions involving the formation of intermolecular beta-sheets.
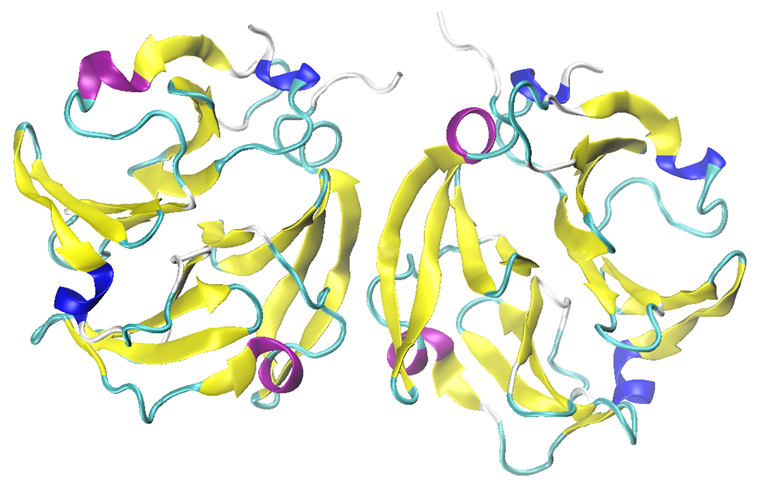
Figure 5. The hemopexin-like domain of MMP-9 in dimeric form (PDB: 1ITV). Image Credit: RedShiftBio
Conclusions
This study examined the proenzymes of three MMPs, MMP-2, -8, and -9, utilizing two MMS systems: the Apollo and the Aurora. This marks the first side-by-side comparison of data obtained from these two systems.
Although all three proteins belong to the same MMP family, and both MMP-2 and MMP-9 are gelatinases, these MMS findings revealed distinct secondary structures in these three proteins. Interestingly, contrary to what their activity would suggest, the structures of MMP-2 and MMP-9 are not more similar to each other than MMP-8.
MMP-9 displays the most intermolecular beta-sheet structure and offers insights into the hemopexin-like dimers. The detected structural difference in the proenzymes matches their respective crystalline structures.
These results offer valuable information about the structural relationship between the proenzymes and active enzymes of MMPs under formulation conditions.

 Download a Brochure
Download a Brochure
References and Further Reading
- Verma, R. P., & Hansch, C. (2007). Matrix metalloproteinases (MMPs): Chemical–biological functions and (Q) SARs. Bioorganic & medicinal chemistry, 15(6), 2223-2268.
- D Avila-Mesquita, C., et al. MMP-2 and MMP-9 levels in plasma are altered and associated with mortality in COVID-19 patients. Biomed Pharmacotherapy. 2021;142:112067.
- da Silva-Neto, P. V., et al. Matrix Metalloproteinases on Severe COVID-19 Lung Disease Pathogenesis: Cooperative Actions of MMP-8/MMP-2 Axis on Immune Response through HLA-G Shedding and Oxidative Stress. Biomolecules. 2022;12(5):604.
- Gelzo, M. et al. Matrix metalloproteinases (MMP) 3 and 9 as biomarkers of severity in COVID-19 patients. Sci Rep 12, 1212 (2022).
- Kendrick, Brent S. et al. "Determining spectroscopic quantitation limits for misfolded structures." Journal of pharmaceutical sciences 109.1 (2020): 933-936.
- Morgunova, E., Tuuttila, A., Bergmann, U., Isupov, M., Lindqvist, Y., Schneider, G., & Tryggvason, K. (1999). Structure of human pro-matrix metalloproteinase-2: activation mechanism revealed. Science, 284(5420), 1667-1670.
- Bertini, I., Calderone, V., Fragai, M., Luchinat, C., Maletta, M., & Yeo, K. J. (2006). Snapshots of the reaction mechanism of matrix metalloproteinases. Angewandte Chemie International Edition, 45(47), 7952-7955.
- Elkins, P. A., Ho, Y. S., Smith, W. W., Janson, C. A., D'Alessio, K. J., McQueney, M. S., ... & Romanic, A. M. (2002). Structure of the C-terminally truncated human ProMMP9, a gelatin-binding matrix metalloproteinase. Acta Crystallographica Section D: Biological Crystallography, 58(7), 1182-1192.
About RedShiftBio
RedShiftBio is redefining the possibilities for analyzing protein structure and concentration.
RedShiftBio has developed a proprietary life sciences platform combining our Microfluidic Modulation Spectroscopy (MMS) and expertise in high-powered quantum cascade lasers that provide ultra-sensitive and ultra-precise measurements of molecular structure. These structural changes affect critical quality attributes governing the safety, efficacy, and stability of biomolecules and their raw materials. This combination of technologies is available to researchers in our fully-automated Aurora and Apollo systems and is backed by a global network of sales, applications, service, and support teams to address all market needs.
Alongside our commitment to further innovation in the formulations and development space, RedShiftBio also supports biopharmaceutical manufacturing with HaLCon, our bioprocess analytics platform, purpose-built to measure protein titer at time of need.
Led by an experienced management team with a proven track record of success in both large instrumentation companies and commercializing disruptive technologies, RedShiftBio is here to support your research, development, and manufacturing goals. Our instruments can be found in the majority of the leading biopharmaceutical companies and CDMOs in the world. We also run product demonstrations and process samples in the StructIR Lab, located in our Boxborough, MA headquarters, as well as at partner sites including the Wood Centre in Oxford, UK, Spectralys/UCB in Brussels, Belgium, and at Sciex laboratories in Redwood Shores, CA.
RedShiftBio is backed by Waters Corporation, Illumina Ventures, Technology Venture Partners, and one undisclosed leading life science company.
Sponsored Content Policy: AZO Life Sciences publishes articles and related content that may be derived from sources where we have existing commercial relationships, provided such content adds value to the core editorial ethos of AZO Life Sciences which is to educate and inform site visitors interested in medical research, science, medical devices and treatments.