The kinetic states of molecules have a vital role in the physiological and metabolic processes in which they participate in biophysics. The amounts of energy, entropy, and enthalpy of protein folding are now specified for the first time in an article published in the journal Proceedings of the National Academy of Sciences (PNAS).
 A team of the Faculty of Physics and the Institute of Nanoscience and Nanotechnology of the UB (IN2UB) introduces for the first time a temperature monitor in optical tweezers to determine the entropy and enthalpy of protein formation. Image Credit: University of Barcelona.
A team of the Faculty of Physics and the Institute of Nanoscience and Nanotechnology of the UB (IN2UB) introduces for the first time a temperature monitor in optical tweezers to determine the entropy and enthalpy of protein formation. Image Credit: University of Barcelona.
To achieve so, the researchers employed a device with optical tweezers that allows them to change the temperature of the experiment between 5 ºC and 40 ºC.
Professor Fèlix Ritort of the UB’s Faculty of Physics and Institute of Nanosciences and Nanotechnology led the research (IN2UB). Its original author is UB researcher Marc Rico-Pasto, and it involves teams from the University of Padova (Italy), the Institute of Bioengineering in Lausanne (Switzerland), and SpliceBio, which has its headquarters in the Barcelona Science Park (PCB).
Optical tweezers to unravel the complexity of living matter
Innovative tools like optical and magnetic tweezers have transformed biophysics research, particularly the study of thermodynamic characteristics in macromolecules like proteins and nucleic acids. Individual molecules may be manipulated with nanometer accuracy (10-9 meters) using forces in the piconewton range with this technique (10-12 newtons).
As a result, researchers can now quantify the thermodynamic characteristics of complex biomolecules at a previously unheard-of level of detail. The use of such methodologies opens up new situations for experimental investigations in the area of thermodynamics using a statistical approach, allowing thermodynamics to be interpreted in ways that were previously only available from a theoretical viewpoint.
Moreover, these methodologies have shortcomings that make it difficult for researchers to distinguish between the sources of the observed forces. Combining various strategies to increase the set of control parameters is now a difficulty in biophysics. This is exactly what the research team did: they included a temperature monitor into the optical tweezers to calculate the entropy and enthalpy of protein folding for the first time.
Energy landscapes in protein folding
Different kinetic states occur during the folding process of cellular macromolecules, ranging from the native state to the denatured state. Transition states, molecular intermediates, and misfolded structures are examples of transitory states that make thermodynamical properties more difficult in experiments with a large number of molecules analyzed concurrently—from the 1023 molecule order, also called the Avogadro’s number.
Transition states are particularly significant to protein folding because of their exceedingly short duration.
Our results reveal that, during the transition state, the protein skeletal structure is already built. However, most of the van der Waals interactions—weak forces—among the residues are not stabilized.”
Fèlix Ritort, Professor and Member, Department of Condensed Matter Physics, University of Barcelona
“Conclusions show that protein folding can be understood as a process defined by two steps. In the first one, the protein reaches the transition state in which the native skeletal structure is built, and water is expelled from the inside of the polypeptide chain. In the second step, the protein collapses, the interactions between protein residues are stabilized, and the protein reaches the native state,” Ritort concludes.
An initial examination of the data indicates that there is a shift in enthalpy and entropy during the transition state that accounts for around 20% of the total enthalpy and entropy observed during the folding.
This phenomenon shows that the protein skeletal structure requires 20% of the interactions between residues. The reading we make from the protein folding goes in line with the most recent hypotheses in the field of protein folding.”
Marc Rico-Pasto, Member, Department of Condensed Matter Physics, University of Barcelona
Given the fact that the protein skeletal structure is formed during the transition stage, scientists claim that they are unable to estimate the number of natural interactions present.
“We can make the first estimation—they say—, but quantifying this result requires some experimental variable that allows us to measure or identify the number of bonds built during the molecular folding in real-time,” explained Rico-Pasto.
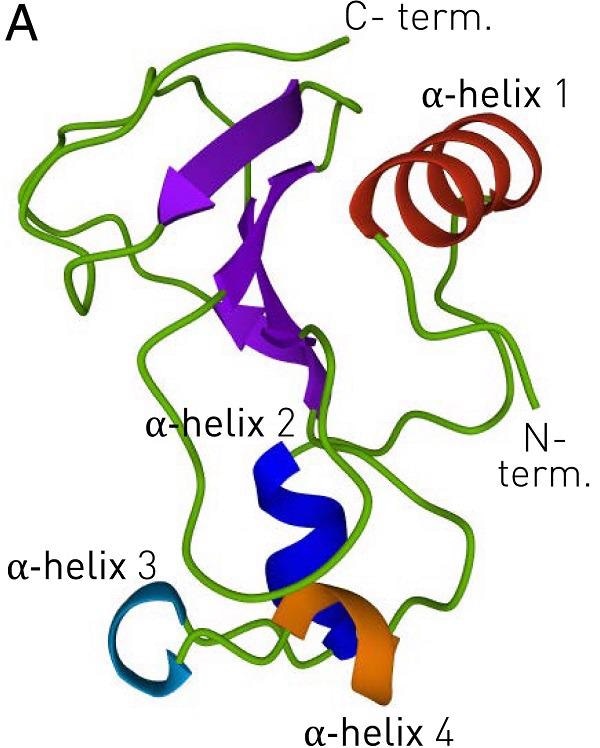
Professor Fèlix Ritort, the head of the Faculty of Physics’ Small Biosystems Lab, led a team that made major advances to the study of thermodynamic characteristics of complex systems in biomolecules. In prior research, the scientists exploited a transition state to extract the barnase protein, a globular biomolecule released by Bacillus amyloliquefaciens.
The barnase is also the reference model for the characterization approach of transition states during the protein folding process, as it does not display intermediate states with a lifespan of more than a millisecond during folding (phi-value analysis).
Source:
Journal reference:
Rico-Pasto, M., et al. (2022) Molten globule–like transition state of protein barnase measured with calorimetric force spectroscopy. Proceedings of the National Academy of Sciences. doi.org/10.1073/pnas.2112382119.