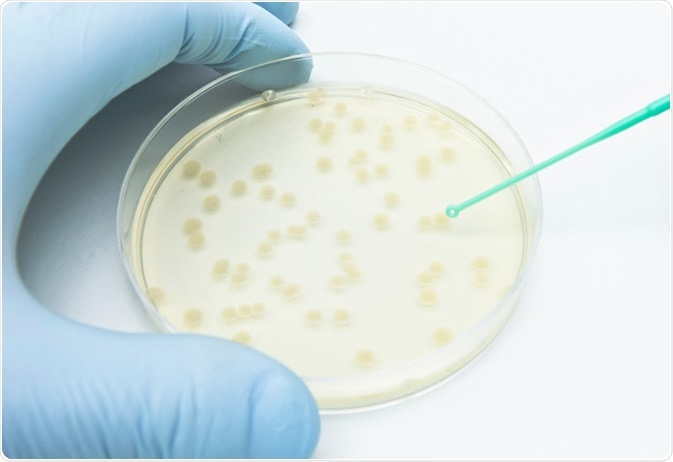
A researcher picking up bacterial colonies on the agar plate using a disposable loop. Image Credit: unoL/Shutterstock.com
How do you look at Microbial Diversity?
The main obstacle in microbiology had been the fact that it is not always possible to grow microorganisms in laboratory conditions – in fact, it is thought that the majority of microorganisms which exist are unculturable. This leads to a bias when examining environmental microorganisms in the laboratory, where a group of organisms will rapidly grow, making it appear highly abundant, but others will not be seen as they are not able to grow.
There was, therefore, a need to develop new techniques to study these environmental microorganisms. This led to the development of “metagenomics”, the study of microorganisms in an environment by analyzing the DNA (genomes) from that environment. The basic principle of metagenomics is the extraction of DNA from an environmental sample, then the analysis of this DNA.
Early metagenomic studies for bacteria developed from the analysis of the 16S ribosomal RNA (rRNA) sequence. The 16S sequence in the genome codes for the ribosomal RNA which forms part of the 30S subunit of the bacterial ribosome, and is therefore highly likely to be present in all bacteria. The 16S rRNA sequence can be present multiple times in a bacterial genome, and because its function has been conserved changes in the 16S rRNA sequence are a good indication of evolutionary time. However, the limitation of 16S rRNA sequence analysis is that it only shows what bacteria are present, and does not give information on how their metabolism works, for example.
It has now become possible to sequence more than just the 16S rRNA sequence, and the development of large scale DNA sequencing means that it is now possible to sequence the whole environmental DNA rather than a few key genes and sequences. However, having this information does not necessarily mean that a lot of new discoveries are made; to see which organisms are present in the environmental sample, genomic information needs to exist so that comparisons can be made.
A problem with this approach is that, for obvious reasons, complete bacterial genomes that are available mostly represent species that could be grown as pure cultures in the laboratory. This means that the “database” is biased to some extent towards bacteria that can be grown, and thus some species present in the environmental sample may be completely missed.
How do you Classify Microorganisms? – Particularly Previously Uncharacterized Ones
It is thought that similarities in the genomes of two organisms would lead to a similarity in their phenotypes, and this is often applied to microorganisms. However, several considerations are needed; depending on which genes are being used as markers, the result may be different.
Also, certain genes can be acquired by horizontal gene transfer – this is where genes are passed between microorganisms that are not directly related. This means that if a marker gene was horizontally transferred between two microorganisms it may appear that these two microorganisms are closely related, and this may in fact not be true.
There is an approach, known as “functional genomics”, which aims to look at the structure, function, and regulation of genes and how these genes in the genome interact with each other, and how they influence the phenotype of the cell or organism. Can something similar be used to classify microorganisms? For example, is it possible to “compare” genomes to see how related two genomes are? -i.e. treating each genome like a gene in functional genomics.
A study by Tsai et al. used this approach to assess the relatedness of nearly 8,000 microbial genomes. Here, the authors used a genome comparison tool called “LAST” to compare genomes, and if two genomes were deemed to be highly similar then a homologous coverage ratio (HCR) was calculated to determine how related these two microorganisms are; HRC would be equal to 1 if two genomes are identical.
The authors saw that bacteria and archaea were distantly related. In most cases, they found that these genomes clustered according to their current classification, but there were some overlaps already at the phylum level. This indicates that some microorganisms classified as belonging to different phyla may be much more closely related than previously thought.
Looking further, the authors found some species which clustered differently. An example is the HCR between Mesoplasma and Mycoplasma species, and the HCR between different Mycoplasma species. The highest HCR between Mesoplasma and Mycoplasma species was ~0.393, but between different Mycoplasma species, the lowest HCR was only ~0.025. This means that in some cases there was less variation between the two different species than within Mycoplasma species. This highlights the possibility that microorganisms are misclassified, and how new, more accurate information may be gained by comparing whole genomes rather than looking at a selected panel of marker genes.
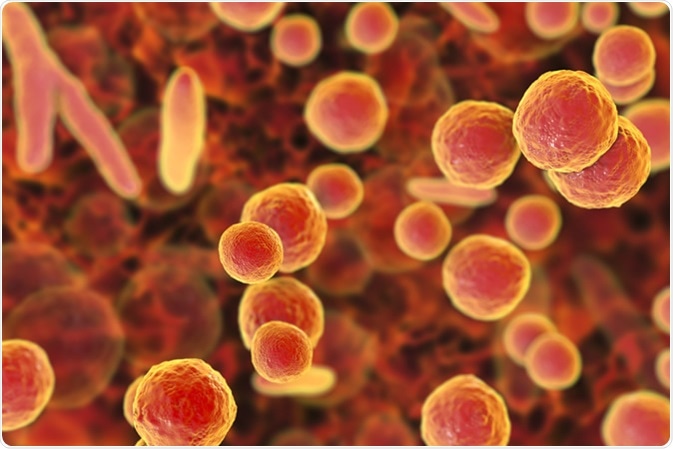
Mycoplasma bacteria, 3D illustration showing small polymorphic bacteria which cause pneumonia, genital and urinary infections - Illustration Credit: Kateryna Kon / Shutterstock
Sources
- Handelsman, J., Metagenomics: Application of Genomics to Uncultured Microorganisms. Microbiol Mol Biol Rev 2004, 68 (4), 669–685. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC539003/
- Janda, J. M., and Abbott, S. L., 16S rRNA Gene Sequencing for Bacterial Identification in the Diagnostic Laboratory: Pluses, Perils and Pitfalls. Journal of Clinical Microbiology 2007, 45 (9), 2761-2764.
- Hugenholtz, P., Exploring prokaryotic diversity in the genomic era. Genome Biology 2002, 3, Article number: reviews0003.1. genomebiology.biomedcentral.com/.../gb-2002-3-2-reviews0003
- sciencedirect.com Horizontal Gene Transfer www.sciencedirect.com/topics/neuroscience/horizontal-gene-transfer
- sciencedirect.com Functional Genomics www.sciencedirect.com/.../functional-genomics
- Tsai, M-H et al., A New Genome-to-Genome Comparison Approach for Large-Scale Revisiting of Current Microbial Taxonomy. Microorganisms 2019, 7 (6), 161. https://www.mdpi.com/2076-2607/7/6/161/htm
Further Reading
Last Updated: Jun 5, 2023