Matrix-assisted laser desorption/ionization time-of-flight mass spectrometry (MALDI-TOF MS) is a high throughput mass spectrometer able to characterize proteins based on their unique spectral fingerprint.
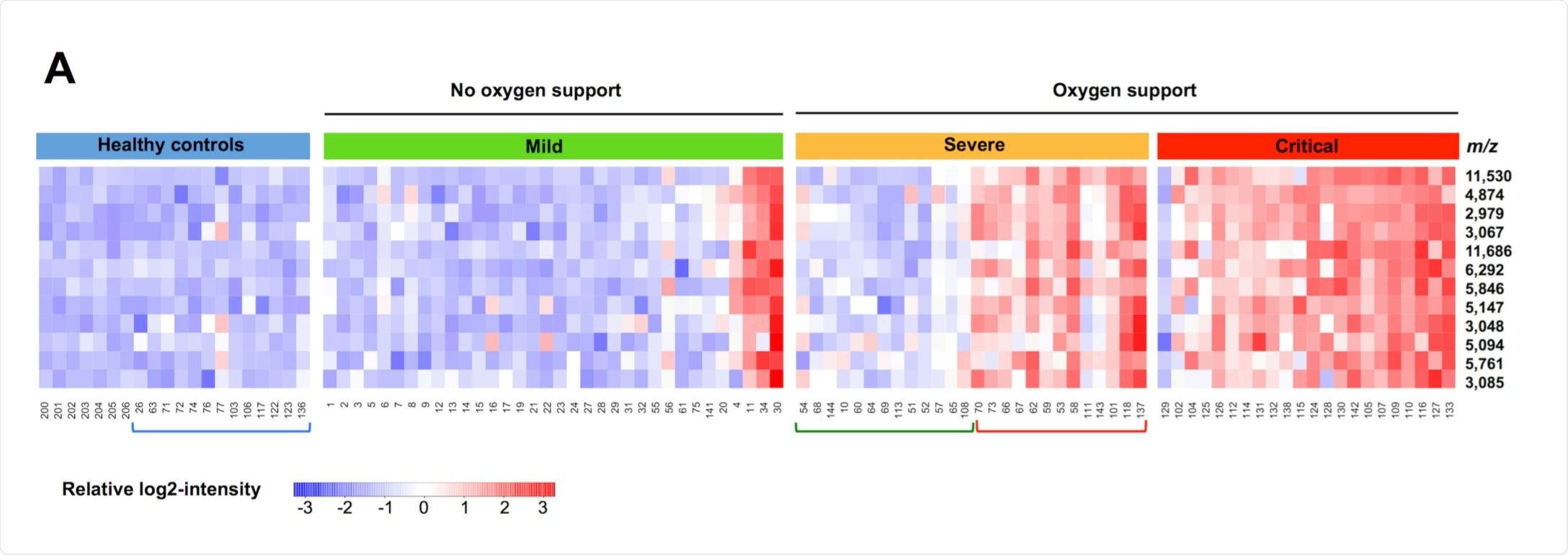
Image Credit: https://www.medrxiv.org/content/10.1101/2020.10.30.20223057v1.full.pdf

 *Important notice: medRxiv publishes preliminary scientific reports that are not peer-reviewed and, therefore, should not be regarded as conclusive, guide clinical practice/health-related behavior, or treated as established information.
*Important notice: medRxiv publishes preliminary scientific reports that are not peer-reviewed and, therefore, should not be regarded as conclusive, guide clinical practice/health-related behavior, or treated as established information.
Gomila et al. recently uploaded a study to the preprint server medRxiv* (November 2020), aiming to utilize the technology to analyze the sera of patients infected with SARS-CoV-2 in a range of disease severity states. They found that their method classed patients into either serious or mild categories with 90% accuracy, and correctly predicted positive outcomes in 85% of cases and negative outcomes in 38% of cases.
Further, the group also utilized liquid chromatography-mass spectrometry to identify five severely upregulated proteins in critical patients, potentially indicating a useful biomarker of disease progression.
Why is this important?
The majority of cases of COVID-19 are asymptomatic or present only mild symptoms, while conversely those that are seriously affected often present exaggerated autoimmune responses, develop viral pneumonia, and require intubation and mechanical intervention.
Therefore, it would be of great advantage to identify key biomarkers that indicate the likely severity of the condition, allowing these patients to be identified early. Current methods are often time-consuming, requiring experienced personnel and specialist equipment. MALDI-TOF MS is comparatively simple and quick to operate and is readily available in most hospitals.
How was the study done?
The group collected samples from 80 SARS-CoV-2 positive patients, confirmed by RT-PCR. They then divided these patients into three categories of severity: mild, for those without or with only mild pneumonia; severe, characterized by dyspnea, a respiratory frequency of fewer than 30 breaths per minute, a blood oxygen saturation of less than 93%, and a PaO2/FiO2 ratio of 50% within 24–48 hours; and critical, for those that exhibited respiratory failure, septic shock, and/or multiple organ dysfunction/failure.
34 were in the mild grouping, 26 severe, and 22 in the critical. Twenty healthy volunteers were used as a control group, 13 of which had previously had and recovered from COVID-19.
The serum samples were purified to obtain the proteins and passed through the MALDI-TOF MS instrument, and data processing techniques applied to identify the 45 most relevant peaks. Separate aliquots of the same samples were additionally used in liquid chromatography-mass spectrometry.
What differences were noted between the groups?
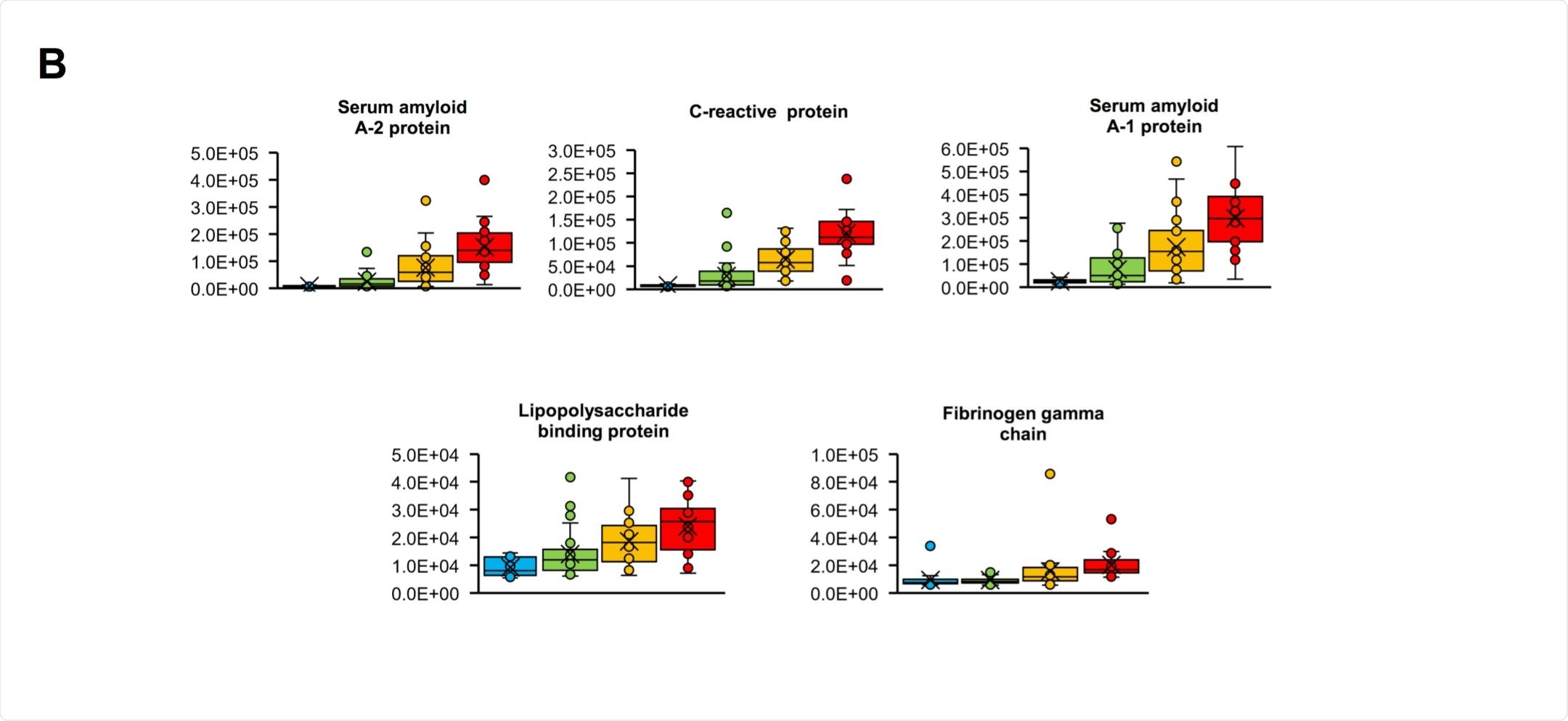
In comparing mild and critical patients, 12 peaks were noted to be at least four times as intense in the critical patients, the five most notable one's likely corresponding to serum amyloid A1, serum amyloid A2, C-reactive, lipopolysaccharide-binding, and fibrinogen gamma chain proteins.
Only two peaks were significantly different between the severe are critical groups, at m/z 11,530 and 11,686, which prompted the additional liquid chromatography-mass spectrometry investigation. The group suggests that the peaks belonged to serum amyloid A1 and serum amyloid A2 proteins, which also showed large increases when comparing mild to severe patients.
Isoforms of serum amyloid A protein have been identified as biomarkers in two other recently published papers and appears that the increased levels may be induced by proinflammatory cytokine interleukin-6, which is secreted by macrophages in response to particular pathogens.
The group noted that those patients in the mild category bore a peptidome signature practically indistinguishable from the healthy controls, which they suggest may be due to the more localized state of infection, with blood serum relatively unaltered at this stage.

 *Important notice: medRxiv publishes preliminary scientific reports that are not peer-reviewed and, therefore, should not be regarded as conclusive, guide clinical practice/health-related behavior, or treated as established information.
*Important notice: medRxiv publishes preliminary scientific reports that are not peer-reviewed and, therefore, should not be regarded as conclusive, guide clinical practice/health-related behavior, or treated as established information.