Cas7-11, the first CRISPR enzyme that can make precise, directed cuts to strands of RNA without hurting cells, was identified and described last year by researchers at MIT’s McGovern Institute for Brain Research.
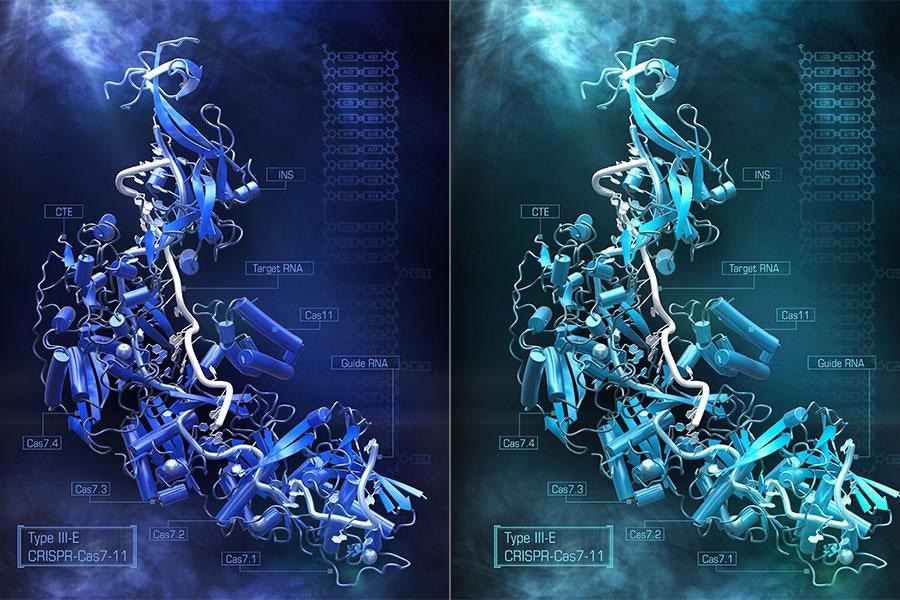 Artistic rendering of the CRISPR Cas 7-11S enzyme. Image Credit: Steve Dixon.
Artistic rendering of the CRISPR Cas 7-11S enzyme. Image Credit: Steve Dixon.
The same research has now demonstrated, in collaboration with partners at the University of Tokyo, that Cas7-11 can be shrunk towards a more compact variant, giving it an even more promising candidate for editing RNA within live cells. The new, compact Cas7-11 enzyme, as well as a comprehensive structural examination of the original enzyme, were published in the journal Cell on May 27th, 2022.
When we looked at the structure, it was clear there were some pieces that weren’t needed, which we could actually remove. This makes the enzyme small enough that it fits into a single viral vector for therapeutic applications.”
Omar Abudayyeh, Research Scientist, McGovern Institute for Brain Research, Massachusetts Institute of Technology
Abudayyeh led the new research along with Jonathan Gootenberg, Research Scientist and McGovern Fellow, and partner Hiroshi Nishimasu from the University of Tokyo.
The scientists, which include former McGovern Institute postdoc Nathan Zhou and University of Tokyo’s Kazuki Kato, regard the new three-dimensional structure of Cas7-11 as a valuable resource for answering concerns about the enzymes’ basic biology and revealing future approaches to tweak their function.
Targeting RNA
CRISPR-Cas9 genome editing technology has allowed researchers the capacity to tweak genes inside human cells during the last decade, which has been a benefit for fundamental research as well as the creation of therapies to cure disease-causing genetic mutations. However, CRISPR-Cas9 only works on DNA, whereas editing RNA is more efficient or beneficial for various scientific and therapeutic objectives.
Any modifications to DNA are generally permanent since a cell preserves its DNA for the rest of its life and transfers an exact replica to daughter cells when it replicates. RNA, on the other hand, is a more transient molecule that is transcribed from DNA and quickly destroyed.
There are lots of positives about being able to permanently change DNA, especially when it comes to treating an inherited genetic disease. But for an infection, an injury, or some other temporary disease, being able to temporarily modify a gene through RNA targeting makes more sense.”
Jonathan Gootenberg, Research Scientist, McGovern Institute for Brain Research, Massachusetts Institute of Technology
The only enzyme that could target RNA had a terrible side effect before Abudayyeh, Gootenberg, and their colleagues identified and defined Cas7-11; when it targeted a specific gene, the enzyme—Cas13—began breaking up all the RNA surrounding it. Cas13’s ability to detect the presence of a piece of RNA makes it beneficial for diagnostic procedures, but not so much for treatments, where targeted cuts are necessary.
Cas7-11 discovery paved the way for a more precise type of RNA editing, similar to the Cas9 enzyme for DNA. The huge Cas7-11 protein, on the other hand, was too enormous to fit within a single viral vector—the empty shell of a virus used by researchers to deliver gene-editing machinery into patients’ cells.
Structural insight
Cryo-electron microscopy, which shoots electron beams on frozen protein samples and evaluates how the beams are transmitted, was used by Abudayyeh, Gootenberg, and Nishimasu to identify the overall structure of Cas7-11. The researchers knew Cas7-11 was made up of five different Cas enzymes fused into a single gene, but researchers did not know how the pieces folded and fit together.
The really fascinating thing about Cas7-11, from a fundamental biology perspective, is that it should be all these separate pieces that come together, but instead you have a fusion into one gene. We really didn’t know what that would look like.”
Jonathan Gootenberg, Research Scientist, McGovern Institute for Brain Research, Massachusetts Institute of Technology
Cas7-11’s structure, captured in the act of engaging both its target tRNA strand as well as the guide RNA that guides that binding, revealed how the components fit together and which portions of the protein were essential for identifying and cutting RNA. This type of structural knowledge is crucial for figuring out how to get Cas7-11 to perform certain tasks within human cells.
The structure also revealed a region of the protein that did not appear to have any function. This discovery suggested that the researchers may eliminate it by re-engineering Cas7-11 to make it smaller while yet retaining its capacity to target RNA.
Abudayyeh and Gootenberg investigated the effects of deleting various parts of this segment, resulting in Cas7-11S, a novel compact form of the protein. Researchers encapsulated Cas7-11S into a single viral vector, transported it into mammalian cells, and successfully targeted RNA using it.
Future research will focus on other proteins that connect with Cas7-11 in the bacteria that it comes from, as well as efforts to exploit Cas7-11 for therapeutic purposes.
“Imagine you could have an RNA gene therapy, and when you take it, it modifies your RNA, but when you stop taking it, that modification stops. This is really just the beginning of enabling that tool set,” Abudayyeh concluded.
Source:
Journal reference:
Kato, K., et al. (2022) Structure and engineering of the type III-E CRISPR-Cas7-11 effector complex. Cell. doi.org/10.1016/j.cell.2022.05.003.