Using machine learning and gene sequencing, scientists from New York University have designed a “developmental atlas” of gene expression in neurons to classify over 250,000 neurons in the brains of fruit flies.
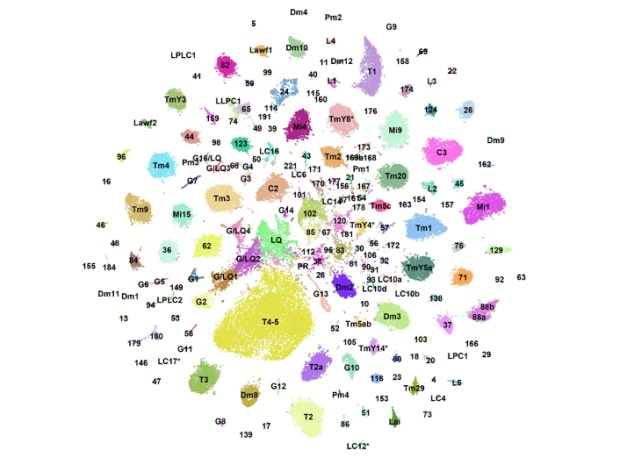
NYU biologists created a “developmental atlas” of gene expression in the neurons of fruit flies. Each dot represents single cells that are organized into color-coded clusters according to their cell type. Image Credit: New York University.
The study, which was recently published in the Nature journal, found that neurons display the most molecular diversity at the time of development and showed a previously unfamiliar type of neurons that is present only before the flies hatch.
Diversity of the different cell types that make up our brains can only be fully understood in light of their developmental history.”
Claude Desplan, Study Senior Author and Biology Professor, New York University
Brains contain countless numbers of different types of neurons. But in spite of sharing the same genetic data, neurons achieve this diversity by switching on different sets of genes in each type of neuron and at each point during their development.
To interpret the diversity of brain cells, scientists have long explored fruit flies, whose brains, despite being relatively simpler than those of human beings, can be utilized as a model system. Scientists have already identified around200 types of neurons and 60,000 cells that make up the optic lobes of fruit flies; optic lobes are the regions of the brain that process visual information, like color vision and detection of motion and objects.
In their latest research reported in the Nature journal, the research team from Desplan’s laboratory sought to fully define the diversity of neurons found in the optic lobe and construct a “developmental atlas” of gene expression, comparing the brain cells of adult flies and studying the variations during development.
The team produced their “atlas” by leveraging a form of a newly developed method, called single-cell mRNA sequencing, which enabled them to trap and sequence mRNA from over 250,000 single cells. Then, using a combination of machine learning techniques, the team assigned all of these cells to a particular type of cell throughout the development
Our datasets almost completely account for the known neuronal diversity of the optic lobes and can serve as a paradigm to understand brain development across species. The ‘atlas’ constitutes an enormous resource for the research community: we can now simply look up whether a particular gene is active or not in any cell type of our choice and at any point during its development.”
Neset Özel, Study Lead Author and Postdoctoral Associate, New York University
The researchers made many discoveries while constructing their “developmental atlas.” Firstly, they identified an entirely new type of neurons in fruit flies that exists only on the surface of the optic lobe at the time of development but is removed via programmed cell apoptosis before the flies hatch.
While we do not yet understand the functions of these previously unknown neurons, neurons with very similar properties—called Cajal-Retzius cells—also exist in mammalian brains, and they are critical for proper brain development.”
Felix Simon, Study Lead Author and Biology Doctoral Student, New York University
The team also discovered that the neurons display the highest concentrations of molecular diversity at the time of development when compared to the adult neurons, enabling cells during development to create connections with particular partner cells—and evade the wrong ones. Consequently, neurons can gain discrete functions and features only because of their developmental history, although their physiological traits in adult brains may be identical.
Özel added, “This has large implications for the studies of neurodevelopmental disorders. Disruptions to neural circuit function could occur entirely due to defects in certain genetic programs that are only transiently active during development and would be impossible to understand by simply looking at the end result.”
Last but not the least, the study demonstrated that neurons that appear identical in form can express diverse sets of genes in the upper part of the brain versus the lower part of the organ. Such differences can give an ability to the flies to carry out different calculations on the visual information received by them—for example, the ground versus the sky.
Source:
Journal reference:
Özel, M. N., et al. (2020) Neuronal diversity and convergence in a visual system developmental atlas. Nature. doi.org/10.1038/s41586-020-2879-3.