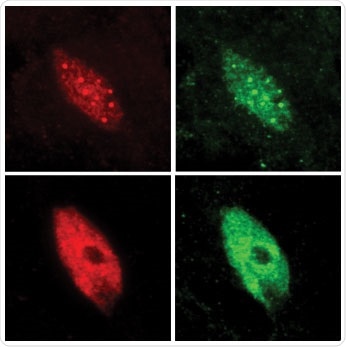
Nuclei of plant cells are seen before and after the plant was exposed to shade. In the top images (before), the transcription factor PIF7 (red) is confined in speckles that contain the plant’s light sensors (green). In the lower images (after), in the shaded plant, PIF7 is released, which is then free to bind to DNA and initiate gene activity. Image Credit: Chan Yul Yoo, Meng Chen lab, UC Riverside.
These results shed new light on how to boost yield and protect the global world food production as climate change reduces the arable land of the planet.
This paper shows, in high resolution, how plants respond to subtle environmental changes on the cellular level. Work that reveals how plants can adapt to greater environmental stresses will be critical as the effects of climate change intensify.”
Joanne Chory, Study Co-Corresponding Author and Director, Plant Molecular and Cellular Biology Laboratory, Salk Institute
Chory is also an investigator from the Howard Hughes Medical Institute and holds the Howard H. and Maryam R. Newman Chair in Plant Biology.
Plants grow faster and taller in the shade in an attempt to break through the canopy and absorb more light. But at the same time, the shaded growing conditions cause these plants to produce seeds or flowers earlier than normal to outpace other plants.
Such responses may benefit wildflowers that grow in a meadow, but on farms, the same kinds of responses can result in decreased production and lead to low-quality and bitter crops—a fact that can be vouched by any gardener whose lettuce has bolted.
In the latest study, the researchers focused on the role of specific transcription factors in triggering the growth response. Transcription factors are essentially proteins that switch the genes on or off by attaching to DNA.
Mutant seedlings that lack transcription factors, known as Phytochrome-Interacting Factors (PIFs) were used for the research work. When these plants were allowed to grow in a shade-stimulating environment, plants lacking certain PIFs did not speed up or elongate their growth but rather continued to grow usually as if they were in full sunlight. The Chory laboratory has already demonstrated the crucial role played by PIF7 in the regulation of shade-induced growth.
Subsequently, the team closely analyzed the role of histones in the process, particularly the histone variant H2A.Z. Histones are essentially proteins that serve like spools for DNA strands. Histones, when modified or exchanged, work to suppress o activate specific genes.
The researchers discovered that canopy shade triggered the removal of the histone H2A.Z at growth-regulating genes via the DNA binding of PIF7, which consequently triggered their expression.
By applying very short intervals to the experiments, the team observed that PIF7 gets activated, attaches its target genes, and riggers the removal of H2A.Z all in less than the first five minutes of the plant experiencing canopy shade.
Our study describes another step towards a mechanistic understanding of how plants alter their gene expression in response to a changing environment.”
Joseph Ecker, Study Co-Corresponding Author and Professor, Genomic Analysis Laboratory, Salk Institute
Joseph Ecker is also an investigator from the Howard Hughes Medical Institute.
Previous analyses had identified the crucial roles of PIFs and H2A.Z in the responses of plants subjected to higher temperatures. Björn Willige, the study co-author and a research specialist in the Chory lab at the Howard Hughes Medical Institute noted that the timing of events was unknown.
Our study reveals the mechanism in close detail and also shows the rapid nature of the response. We found that when PIF7 is active, it binds to DNA. And our data indicate that this leads to the removal of H2A.Z from the DNA. Subsequently, genes are activated, and then this induces growth, to outcompete the neighboring plants.”
Björn Willige, Study Co-Author, Research Specialist, Howard Hughes Medical Institute
According to Mark Zander, the study co-author and an assistant professor from the Waksman Institute of Microbiology at Rutgers University, the pace of the process was rather unexpected. He observed that apart from activating the response in less than five minutes, the histone landscape also recovered rapidly once the shade was removed.
“When we removed shade, the levels of H2A.Z at PIF7 target genes went back to normal within 30 minutes. I was surprised by how dynamic the process is, which is the foundation for the elegance of our study,” added Zander.
PIFs play a major role in the development, growth, and pest defense of plants. The researchers are therefore hoping that their results can be translated to other similar plant responses that are significant for farmers, particularly with respect to aiding plants to be more resistant to climate change.
The Harnessing Plants Initiative of the Salk Institute is seeking to help solve climate change by improving the natural ability of plants to capture and preserve carbon.
Source:
Journal reference:
Willige, B. C., et al. (2021) PHYTOCHROME-INTERACTING FACTORs trigger environmentally responsive chromatin dynamics in plants. Nature Genetics. doi.org/10.1038/s41588-021-00882-3.