The so-called plant microbiome—the microorganisms in a plant and its immediate environment—impacts plant health, survival, and fitness. This was demonstrated in a recent study performed.
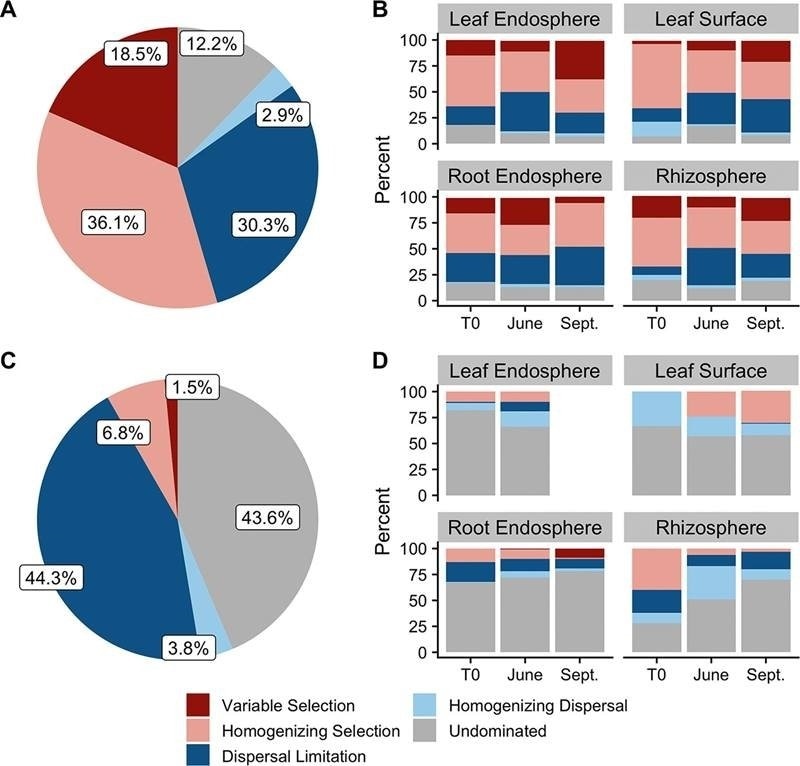
At left, overall dominance of microbiome assembly processes (see legend) for archaea and bacteria (A) and fungi (C). Right, relative dominance of same assembly processes in each habitat-sample date combination for archaea and bacteria (B) and fungi (D). Image Credit: Copyright Nicholas Dove, Oak Ridge National Laboratory, from Dove, N. C., et al., Assembly of the Populus microbiome is temporally dynamic and determined by selective and stochastic factors.
The initial assembly of the microbiome is considered to be specifically significant. Assembly relates to the processes that exhibit the potential to generate a number of variety of species inside the microbiome. This study characterized the initial assembly of the microbiome in numerous types of poplar trees. The study discovered that there was a considerable alteration in the composition of the microbiome over time.
For bacteria and archaea present in the microbiome, the passage of time made the amount of variation reduce. But, alteration among fungi in the microbiome was molded with the help of various processes. The genetic makeup of poplar trees was considered to be less of a factor compared to what the scientists had anticipated.
The impact
The initial assembly of a plant microbiome might help set the future of the microbiome. This can identify the complete future health of the plant. But while researchers have a good understanding of the initial microbiome assembly of agricultural crops and grasses, they know less regarding the initial microbiome of long-lived trees, like poplar.
Poplar trees might be considered as outstanding candidates for biofuels as well as other applications. If researchers gain a better insight into the microbiome of the plants, these trees could be of greater use to them. For instance, the results of this new study could help researchers utilize microbes to enhance the growth and health of popular trees.
Summary
The study performed in recent times has demonstrated that the plant microbiome, especially the initial assembly of this microbiome, tends to impact plant health, survival, and fitness. In this study, researchers marked the initial assembly of Populus microbial communities throughout ten genotypes attributed to two poplar species in a common garden.
The microbiome was sequenced by the scientists inside the leaf surface, leaf and root tissues (leaf and root endosphere), and root surface and its immediate surroundings (rhizosphere). They coupled molecular analyses with source tracking models and ecological assembly models to explain the assembly of the Populus microbiome in the course of the first growing season.
The researchers discovered that the composition of the microbiome altered considerably over time throughout all plant-associated habitats and host genotypes. Considering bacteria and archaea, these alterations were dominated by strong homogenizing selection (accounting for 29% to 62% of pairwise comparisons).
But, the fungal assembly was normally characterized by several ecological assembly processes (a mixture of weak selective and dispersal processes). Intriguingly, genotype, while a considerable moderator of microbiome composition, normally described less variation compared to sample date throughout plant-associated habitats.
The scientists outlined a set of core genera that accounted for 36% of the microbiome typically. The comparative abundance of this core community was persistent over time. Besides, with the help of source-tracking modeling, they identified that new microbial taxa colonize from both aboveground and belowground sources.
Having integrated with the ecological assembly null models, they discovered that both selective and dispersal processes described the variations between exospheric (that is., rhizosphere and leaf surface) and endospheric microbiomes. Altogether, the findings indicate that the Populus microbiome’s initial assembly is time-, genotype-, and habitat-dependent and is driven by both stochastic and selective factors.
Source:
Journal reference:
Dove N C., et al. (2021) Assembly of the Populus Microbiome Is Temporally Dynamic and Determined by Selective and Stochastic Factors. mSphere. doi.org/10.1128/msphere.01316-20.