At times, the molecular machinery makes alterations to the wrong part of a host’s genome, generating the likelihood of an effort to repair a genetic mutation in one location of the genome can create a harmful new mutation in the other accidentally. This is considered to be one of the huge challenges faced while using CRISPR-based gene editing on humans.
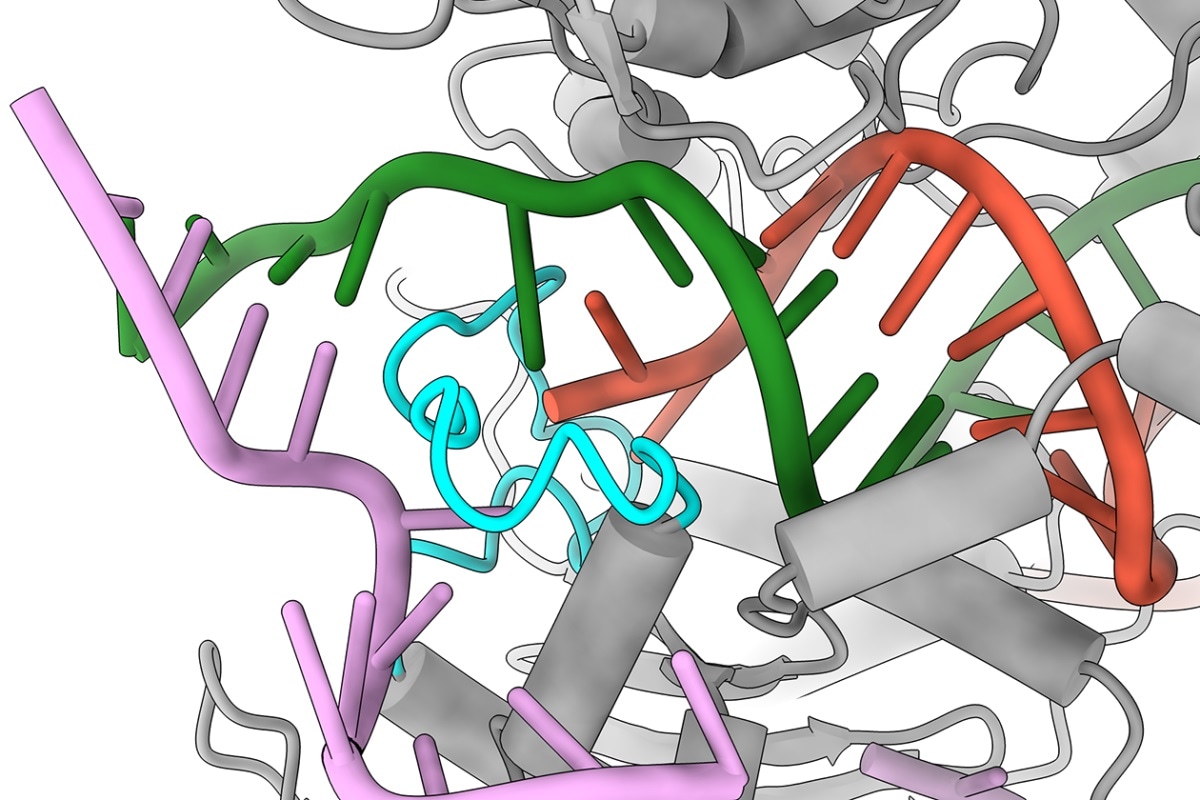
UT Austin researchers were surprised to discover that when Cas9 encounters a mismatch in a certain part of the DNA (red and green), instead of giving up and moving on, it has a finger-like structure (cyan) that swoops in and holds on to the DNA, making it act as if it were the correct sequence. Image Credit: Jack Bravo/University of Texas at Austin.
Now, however, researchers at The University of Texas at Austin have redeveloped a vital component of a commonly used CRISPR-based gene-editing tool, known as Cas9, to be thousands of times less probable to attack the wrong stretch of DNA when being just as successful as the original version, making it possibly much safer. The study is published in the journal Nature.
“This really could be a game changer in terms of a wider application of the CRISPR Cas systems in gene editing,” stated Kenneth Johnson, the study’s co-senior author and professor, of Molecular Biosciences and David Taylor is a co-senior author of the study and an assistant professor of molecular biosciences. The co-first authors of the paper are postdoctoral fellows Jack Bravo and Mu-Sen Liu.
Other laboratories have restructured Cas9 to decrease off-target interactions, but until now, all these forms enhance precision by sacrificing speed. SuperFi-Cas9, a dubbed new version, is 4,000 times less probable to cut off-target locations but just as quickly as naturally occurring Cas9.
Bravo says that the various lab-produced versions of Cas9 can be thought of as different models of self-driving cars. The majority of models are indeed safe, but they have the highest speed of 10 miles per hour.
They’re safer than the naturally occurring Cas9, but it comes at a big cost: They’re going extremely slowly. SuperFi-Cas9 is like a self-driving car that has been engineered to be extremely safe, but it can still go at full speed.”
Jack Bravo, Postdoctoral Fellow, The University of Texas at Austin
Until now, scientists have established the usage of SuperFi-Cas9 on DNA in test tubes. They are now joining hands with other scientists who intend to verify SuperFi-Cas9 for gene editing in living cells. They are also working to build safer and more active versions of Cas9.
CRISPR-based gene-editing instruments are adapted from naturally occurring systems in bacteria. A Cas9 protein hovers around in the surroundings, in nature, looking for DNA with a very particular sequence of 20 letters, such as the X on a pirate map indicating “dig here.”
At times, when most of the letters are right, excluding those in spots 18 through 20, Cas9 will still go ahead and dig in. This is known as a mismatch, and it could have catastrophic results in gene editing.
A technique called kinetics-guided structure determination was developed by Taylor and Johnson. It used a cryo-electron microscope to take photographs of Cas9 in action while it interacted with the mismatched DNA. This was done in the Sauer Structural Biology Lab.
The researchers were surprised to find that while Cas9 encounters this kind of mismatch in spots 18 through 20, rather than giving up and moving on, it comprises a finger-like structure that dives in and gets hold of the DNA, letting it act as if it were the right sequence.
Typically, the DNA is left a bit floppy by a mismatch; however, this finger-like structure balances it.
It’s like if you had a chair and one of the legs was snapped off and you just duct taped it together again. It could still function as a chair, but it might be a bit wobbly. It’s a pretty dirty fix.”
Jack Bravo, Postdoctoral Fellow, The University of Texas at Austin
Cas9 does not take the other steps required to cut the DNA and do edits without that extra stability in the DNA. None of them had ever noticed this added finger doing such stabilization earlier.
“This was something that I could never have, in a million years, imagined in my mind would have happened,” Taylor stated.
Depending on this understanding, they restructured the extra finger on Cas9. Hence, rather than stabilizing the DNA part that contained the mismatch, the finger is instead set aside from the DNA, preventing Cas9 from continuing the cutting and editing process of the DNA. The result is SuperFi-Cas9, which is a protein that cuts the appropriate target just as freely as naturally occurring Cas9. However, it is much unlikely to cut the incorrect target.
Source:
Journal reference:
Bravo, J. P. K., et al. (2022) Structural basis for mismatch surveillance by CRISPR–Cas9. Nature. doi.org/10.1038/s41586-022-04470-1.