Reviewed by Danielle Ellis, B.Sc.May 30 2022
An international collaboration of scientists has discovered a new technique to kill cancers that are difficult to treat, according to an article published in Nature. Current immunotherapies, including Nobel Prize-winning check-point blocking antibodies, are ineffective against these tumors.
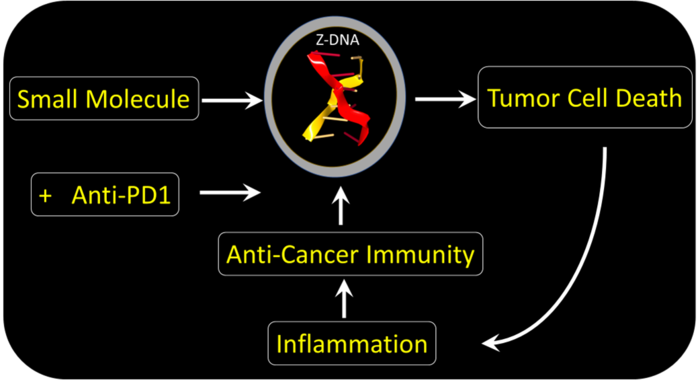 A small molecule induces Z-DNA formation in tumors to activate an inflammatory cell-death pathway that is expressed in the cells supporting growth of the cancer cell. The resulting immune response is specific for the mutant proteins expressed by cancer cells. Tumor killing is further enhanced by infusing an anti-PD-1 antibody that neutralizes the cancer cell’s ability to inhibit the responding immune cells. Image Credit: Alan Herbert.
A small molecule induces Z-DNA formation in tumors to activate an inflammatory cell-death pathway that is expressed in the cells supporting growth of the cancer cell. The resulting immune response is specific for the mutant proteins expressed by cancer cells. Tumor killing is further enhanced by infusing an anti-PD-1 antibody that neutralizes the cancer cell’s ability to inhibit the responding immune cells. Image Credit: Alan Herbert.
Z-DNA is used in this method. It has a left-handed twist, unlike Watson and Crick B-DNA, which twists to the right. Z-DNA regulates the immune response to viruses, for example. Two proteins that particularly detect Z-DNA, AADR1, and ZBP1, are involved in the reaction.
They achieve it by using a Zα domain that has a high affinity for the Z-DNA structure. Dr Alan Herbert of InsideOutBio, a collaborating author on the study, was the first to discover the Zα domain.
As previously demonstrated by Dr Sid Balachandran, the other communication author on the study, the ADAR1 Zα domain switches off the immune response to self, whereas the other ZBP1 Zα domain switches on pathways that destroy virally infected cells. ADAR1 and ZBP1 interactions decide whether a tumor cell lives or dies.
Interferon activates both Zα proteins during inflammation. Zα Proteins are not generally seen in healthy cells. Both proteins are found in tumors, particularly in normal cells known as fibroblasts, which cancer cells use to fuel their development. ADAR1 is normally used by cancers to inhibit cell death pathways that would otherwise destroy the tumor.
The researchers discovered a small molecule that bypassed ADAR1 inhibition and induced tumor cell death via ZBP1. The drug works independently of the cancer-causing mutation. The type of cell death that is caused is immunogenic.
The immune reaction kills the fibroblasts that help the tumor development. As a result, the drug improves the efficacy of immunotherapy utilizing a PD-1-targeted checkpoint blocking antibody. The drug belongs to the curaxin family and was first used in the clinic for a different cause. In Phase I studies, the chemical was shown to be safe, but further study is needed to confirm that its clinical usage in conjunction with anti-PD1 gives a benefit in the treatment of cancers.
This outcome is the work of a highly collaborative team. It is a nice milestone in our understanding of how alternative DNA conformations, like Z-DNA, play an important role in human biology. The paper shows how basic research can lead to new and unexpected therapies. The process has taken a long time, starting with the initial discovery of the Zα domain and then to the identification of Zα DNA variants that cause genetic diseases in humans.”
Dr Alan Herbert, Study Communicating Author, InsideOutBio
“These discoveries validated a biological role for Z-DNA. The work now has led us to a new therapeutic approach for the treatment of cancer. It is a pleasing turn around given that the initial Z-DNA discoveries were widely dismissed by the scientific community as of little biological importance and further work was not ranked as worthy of funding by the National Institutes of Health peer review panels,” Dr Herbert notes.
Dr Herbert adds, “The paper was only possible through the collaborations of the team at Fox Chase Cancer Center led by Dr Balachandran and the computational scientists at the Higher School of Economics lad by Dr. Poptsova, with help from Dr Walkley at University of Melbourne, Dr Xu form University of Miyazaki and Dr Thomas at St. Jude Children’s Research Hospital. It is a sign of the times that this collaboration succeeded though I have none meet any of the others in person.”
In normal cells, Z-DNA is generated by sequences termed flipons, which may reversibly switch to the left-handed conformation under physiological circumstances. The geometry was discovered by chance in the first X-ray crystallography-solved synthetic crystal.
Other types of flipons exist, and they are likely to serve vital functions in biology as well. Flipons are extremely dynamic structures that have proven difficult to research since determining their precise conformation inside cells is difficult.
It is similar to other highly dynamic systems in physics where you can only find the flipon conformation by making a direct measurement, but only if the act of measurement does not bias the result.”
Dr Alan Herbert, Study Communicating Author, InsideOutBio
InsideOutBio is a privately funded early-stage biotech startup that is developing cancer treatments.
Source:
Journal reference:
Zhang, T., et al. (2022) ADAR1 masks the cancer immunotherapeutic promise of ZBP1-driven necroptosis. Nature. doi.org/10.1038/s41586-022-04753-7.