RNA polymerases are involved in the transcription of genes in all living organisms and are essential to life on earth. While there are some differences between RNA polymerases across the kingdoms of life, many of their basic structures and mechanisms of action have been conserved.
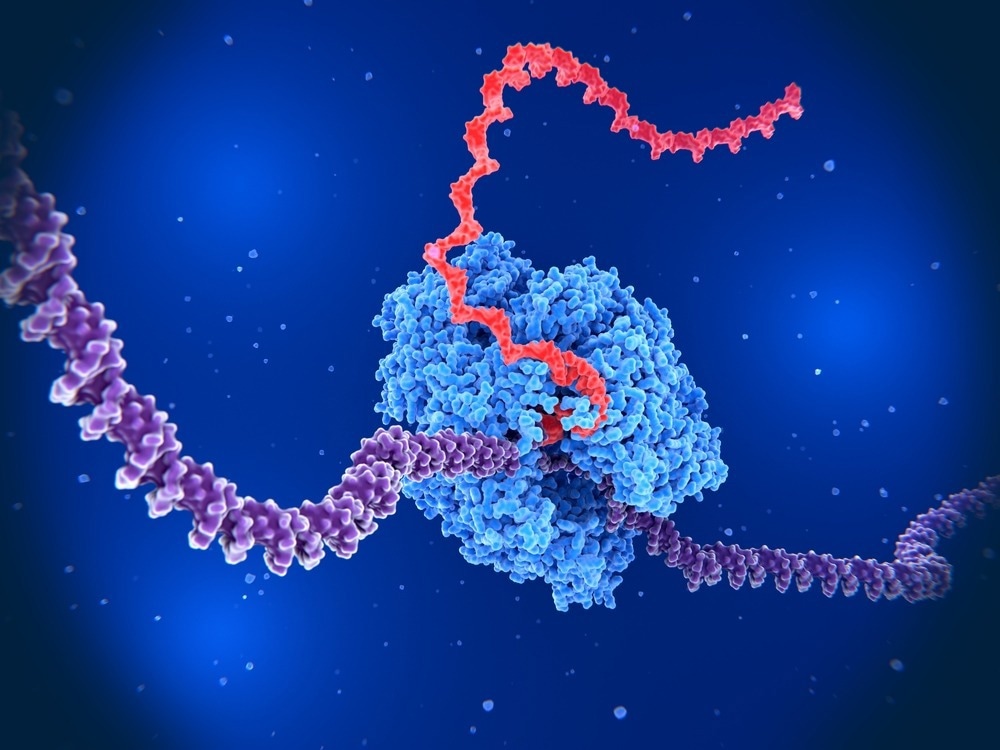
Image Credit: Juan Gaertner/Shutterstock.com
The role of RNA polymerase
RNA polymerase is an enzyme in all living organisms and is involved in gene transcription, in which RNA is synthesized from a DNA template. This is the first step in gene expression and is carried out using similar mechanisms across all six kingdoms of life, from prokaryotic eubacteria and archaea to eukaryotic protists, fungi, plants, and animals.
Transcription is initiated when RNA polymerase binds to a promoter sequence on the DNA. After initiation, elongation can commence, in which the RNA strand is synthesized by adding new nucleotides. The RNA polymerase moves across the strand, adding a complementary RNA nucleotide for each nucleotide present on the DNA strand.
Transcription is complete when the newly formed strand of RNA is released. This RNA will then be translated into proteins or remain as functional non-coding RNA, which plays a role in regulating gene expression.
The structure of RNA polymerases
As well as sharing similarities in their basic mechanisms of action, all of the multisubunit RNA polymerases present today also share a common structural core. This is because they are derived from the RNA polymerase of the last universal common ancestor of eubacteria, archaea, and eukaryotes, which lived roughly 3.5-3.8 billion years ago.
The ancestral RNA polymerase was likely similar in structure to those found in eubacteria today, which have five core subunits. All five of these subunits are highly conserved and present in all living organisms, highlighting that they perform essential functions in the transcription process, allowing for little change or innovation in their structure.
After the separation of eubacteria, the complexity of RNA polymerases increased as additional subunits were added throughout evolution. This increased complexity is linked to archaeal and eukaryotic genomes being stabilized and compacted with histones or histone-like proteins that restrict access to the DNA, meaning additional basal transcription factors are required for transcription to take place.
As a result, archaeal RNA polymerase contains 11-13 subunits depending on the species, many of which are important for stabilizing the interactions between RNA polymerase and the DNA, RNA, and transcription factors.
Transcription in eukaryotic organisms is even more complex than in archaea. Instead of having only one type of RNA polymerase involved in transcription, as seen in prokaryotes, eukaryotes have evolved multiple different types of RNA polymerase.
Eukaryotic RNA polymerases
In protists, fungi, and animals, there are three forms of RNA polymerase: RNA polymerase I, II, and III. All three of these types share a common ancestor and exhibit similarities in subunit organization, with all subunits in RNA polymerase II having equivalent subunits in RNA polymerase I and III.
The number of subunits present in eukaryotic RNA polymerases has increased throughout evolution, from 12 in RNA polymerase II to 14 in RNA polymerase I and 17 in RNA polymerase III. The additional subunits present in I and III are due to the permanent recruitment of transcription factors into the enzyme.
These different forms of RNA polymerase are responsible for the transcription of distinct, non-overlapping subsets of genes. RNA polymerase I synthesize ribosomal RNA, which is used in the production of ribosomes, whereas RNA polymerase II is involved with the transcription of protein-coding genes into messenger RNA. After transcription, messenger RNA carries information from the nucleus to ribosomes, where protein synthesis occurs.
RNA polymerase II also facilitates the production of microRNA, which regulates gene expression. RNA polymerase III helps to produce short, non-coding RNAs such as tRNA and 5S ribosomal RNA, which are essential to protein synthesis and cell functioning.
As well as these three RNA polymerases, plants contain an additional two types, RNA polymerase IV and V. The subunit composition of both types is poorly characterized, although they are known to transcribe small interfering RNA that plays a role in gene silencing, a process that prevents the expression of specific genes where necessary.
Image Credit: Juan Gaertner/Shutterstock.com
What are the real-world applications of our knowledge of RNA polymerases?
Over 50 years have passed since the discovery of the first RNA polymerases, and much has now been uncovered about these enzymes' structure, function, and action.
This research has improved our understanding of gene transcription as well as the mechanisms that lead to the misregulation of gene expression and thereby cause many human diseases. Not only has this knowledge been utilized in developing some anticancer drugs, which target the action of different RNA polymerases, but it has also shown potential for use in the treatment of diseases such as diabetes and immune disorders.
As well as this, the biochemical differences between eukaryotic and bacterial RNA polymerases have been exploited to produce antibiotics that treat bacterial infections such as tuberculosis by targeting the action of bacterial RNA polymerase without affecting human RNA polymerases.
Many of the recent developments in our knowledge of RNA polymerases and the resulting applications of this knowledge can be attributed to advancements in the experimental methods and technologies used in this field of research. Continued utilization of these cutting-edge approaches in future research will help further our understanding of these enzymes and their essential role in the life of all organisms.
From DNA to Protein; The Central Dogma of Molecular Biology
Sources:
- Martin, R.D., Hébert, T.E. and Tanny, J.C. (2020). Therapeutic targeting of the general RNA polymerase II transcription machinery. International Journal of Molecular Sciences, 21(9), p.3354. https://doi.org/10.3390/ijms21093354
- Ferreira, R., Schneekloth Jr, J.S., Panov, K.I., Hannan, K.M. and Hannan, R.D. (2020). Targeting the RNA polymerase I transcription for cancer therapy comes of age. Cells, 9(2), p.266. https://doi.org/10.3390/cells9020266
- Carter, R. and Drouin, G. (2010). The increase in the number of subunits in eukaryotic RNA polymerase III relative to RNA polymerase II is due to the permanent recruitment of general transcription factors. Molecular biology and evolution, 27(5), pp.1035-1043. https://doi.org/10.1093/molbev/msp316
- Reines, D. (2020). Recent advances in understanding RNA polymerase II structure and function. Faculty Reviews, 9. https://doi.org/10.12703/b/9-11
- Werner, F. (2007). Structure and function of archaeal RNA polymerases. Molecular microbiology, 65(6), pp.1395-1404. https://doi.org/10.1111/j.1365-2958.2007.05876.x
- Barba-Aliaga, M., Alepuz, P. and Pérez-Ortín, J.E. (2021). Eukaryotic RNA Polymerases: the many ways to transcribe a gene. Frontiers in Molecular Biosciences, 8, p.663209. https://doi.org/10.3389/fmolb.2021.663209
Further Reading
Last Updated: Nov 11, 2022