As of 2018, 37.9 million people worldwide are known to be living with Human Immunodeficiency Virus (HIV). This is a major epidemic with 1.7 million new infections in 2018 alone.
.jpg)
Image Credit: RAJ CREATIONZS/Shutterstock.com
These high rates of infection provide a large gene pool from which resistant strains can emerge. Although there are two variants of HIV which have overcome the species barrier, the HIV-1 variant is more virulent and widespread and therefore is the variant that will be discussed.
Infection
HIV enters the body via the transmission of infected bodily fluids. After exposure, the virus is capable of entering all cells with CD4 receptors; these include T-cells, macrophages, dendritic cells, and astrocytes.
The envelope glycoprotein on the surface of the HIV-1 virus recognizes the CD4 receptor alongside either CCR5 or CXCR4 receptors, which initiates cell fusion. In particular, HIV has a high affinity for the T-cell CD4 receptor.
Once the virus has fused with the cell it releases the viral capsid into the cytoplasm. This capsid contains two identical RNA strands containing 11 genes, alongside RNA reverse transcriptases and integrase enzymes.
The capsid is taken up by the endosome thereby releasing its contents into the cytoplasm. The RNA reverse transcriptase synthesizes DNA from the viral RNA, this DNA is then transferred into the nucleus, where it can be incorporated into the host genome. Finally, hijacking the host cell’s metabolism to produce the encoded viral proteins and new viral cells.
Infected T-cells have a half-life of 2-4 days. They are killed in one of three ways; by the cytotoxic proteins, the viral DNA produces, cell lysis, or cytotoxic T-cells of the immune system. Not only does the destruction of T-cells reduce the number available in the immune system but the Nef and Tat proteins produce by HIV prevents the formation of new healthy T-cells.
Therefore, long term infection results in immunodeficiency. Even if the pool of infected T-cells is reduced HIV persists in longer-lived cells like macrophages, astrocytes, or memory T cells, increasing the chance of reinfection.
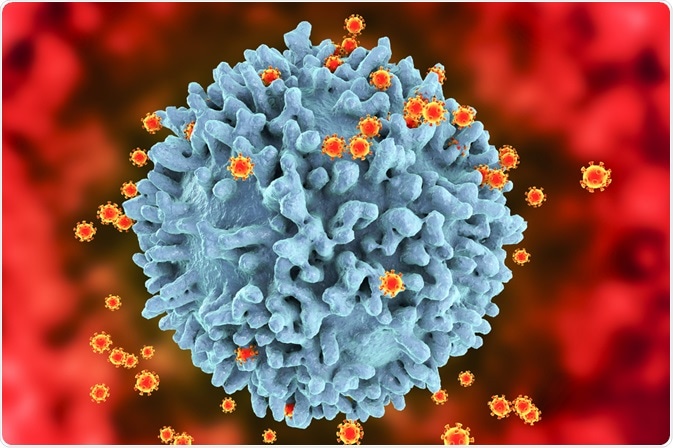
Image Credit: Kateryna Kon/Shutterstock.com
Drug Targets
There are several classes of retroviral drugs currently recommended by the World Health Organisation (WHO). One class of antiretrovirals, CCR5 antagonists, prevents the entry of HIV into the cell by causing allosteric changes in the conformation of CCR5, preventing HIV from binding to the cell.
The other two classes are nucleoside reverse transcriptase inhibitors and non-nucleoside reverse transcriptase inhibitors, target the reverse transcription phase of infection, with modified nucleosides that prevent further elongation and allosteric inhibition of the reverse transcriptase, respectively.
The fourth class of antiretrovirals, integrase strand transfer inhibitors, target the incorporation of the viral DNA into the host’s genome. And the final class, protease inhibitors, target the HIV-1 protease processing of viral proteins.
Resistance
The resistance of HIV derives from the reverse transcription phase. Viral reverse transcriptase is error-prone with an average of one incorrect nucleotide per infection. After only two weeks of infection by a single HIV variant, the number of variants is innumerable.
While this can lead to mutations that are detrimental to the virus, it can also lead to mutations that are beneficial to drug resistance. Each antiretroviral drug has a genetic barrier, this is the number of mutations required to become resistant.
HIV strains that carry drug resistance are often less fit than their wild type counterparts. Consequently, they rarely appear or rise to detectable levels without the selective pressure of antiretrovirals.
However, resistance to one type of antiretroviral does not confer resistance to other types, and whilst resistance can be obtained within a drug type, some drugs increase the susceptibility of HIV to drugs of the same type. Therefore, a comprehensive understanding of drug interactions is crucial to combating resistance.
Overcoming resistance
To contain HIV, the WHO has set a target 90-90-90. That 90% of people know they are HIV positive, of which 90% are being treated, and 90% of them have suppressed viral loads.
They also recommend not using antiretrovirals if the resistance to a drug stands at over 10% of the HIV-positive population in that country. Combining this approach with early recognition of drug resistance within patients and changing their antiretrovirals accordingly.
Most antiretrovirals are sufficiently effective with a high enough genetic barrier for the long-term suppression of HIV-1 replication counterparts. The major issue of resistance comes from incomplete adherence to the drug regiment providing a selection pressure on the HIV virus to develop the resistance mutation.
This resistant variant can then be transmitted to new hosts through primary infection with the resistant strain.
Summary
HIV is a rapidly mutating virus that attacks immune system cells making it difficult to treat. The use of antiretrovirals controls the infection but does not cure it. This exposure to the selective pressure of antiretrovirals leads to resistant HIV populations.
Although new drugs are being developed to overcome this acquired resistance, the current method for control is using a combination of antiretrovirals. Different antiretrovirals target different aspects of HIV infection, therefore an HIV that carries a mutation for the resistance to one type of antiretroviral, will not necessarily be resistant to all antiretroviral types.
Through this method, HIV infection can be kept under control even if it cannot be not cured.
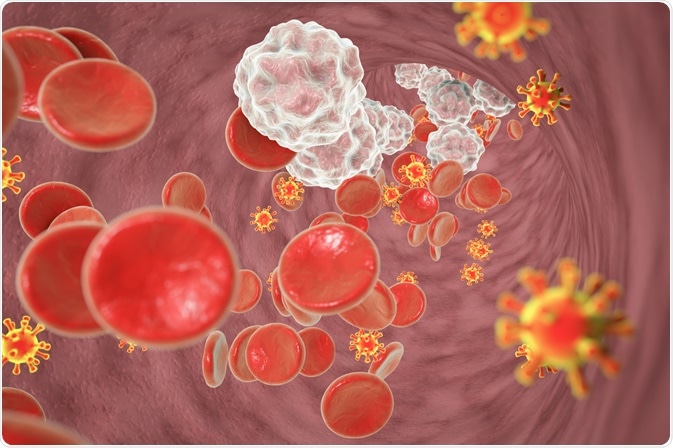
Image Credit: Kateryna Kon/Shutterstock.com
References
- Clutter, D. S. et al. (2016) ‘HIV-1 Drug Resistance and Resistance Testing’, Infection, genetics and evolution: journal of molecular epidemiology and evolutionary genetics in infectious diseases, 46, pp. 292–307. doi: 10.1016/j.meegid.2016.08.031.
- German Advisory Committee Blood (Arbeitskreis Blut) and Subgroup ‘Assessment of Pathogens Transmissible by Blood’ (2016) ‘Human Immunodeficiency Virus (HIV)’, Transfusion Medicine and Hemotherapy, 43, pp. 203–222. doi: 10.1159/000445852.
- Ghosh, A. K., Osswald, H. L. and Prato, G. (2016) ‘Recent Progress in the Development of HIV-1 Protease Inhibitors for the Treatment of HIV/AIDS’, Journal of Medicinal Chemistry, 59(11), pp. 5172–5208. doi: 10.1021/acs.jmedchem.5b01697.
- Klasse, P. J. (2012) ‘The molecular basis of HIV entry’, Cellular Microbiology, 14(8), pp. 1183–1192. doi: 10.1111/j.1462-5822.2012.01812.x.
- Nakata, H. et al. (2019) ‘Activity and structural analysis of GRL-117C: a novel small molecule CCR5 inhibitor active against R5-tropic HIV-1s’, Scientific Reports, 9(1), pp. 1–10. doi: 10.1038/s41598-019-41080-w.
- Tarasova, O., Poroikov, V. and Veselovsky, A. (2018) ‘Molecular Docking Studies of HIV-1 Resistance to Reverse Transcriptase Inhibitors: Mini-Review’, Molecules, 23(5), pp. 11–13. doi: 10.3390/molecules23051233.
- United Nations Joint Programme on HIV/AIDS (UNAIDS) (2019) UNAIDS Data 2019. Available at: https://www.unaids.org/sites/default/files/media_asset/2019-UNAIDS-data_en.pdf (Accessed: 2 April 2020)
- Weber, J. et al. (2013) ‘Resistance Mutations outside the Integrase Coding Region Have an Effect on Human Immunodeficiency Virus Replicative Fitness but Do Not Affect Its Susceptibility to Integrase Strand Transfer Inhibitors’, PLoS ONE, 8(6). doi: 10.1371/journal.pone.0065631.
- World Health Organization (WHO) (2019) HIV Drug Resistance Report. Available at: https://www.who.int/. (Accessed: 28 March 2020)
Last Updated: Sep 12, 2022