For the first time, researchers from the University of Warwick have observed the movements of cell muscles in the form of minute filaments of proteins at unparalleled detail.
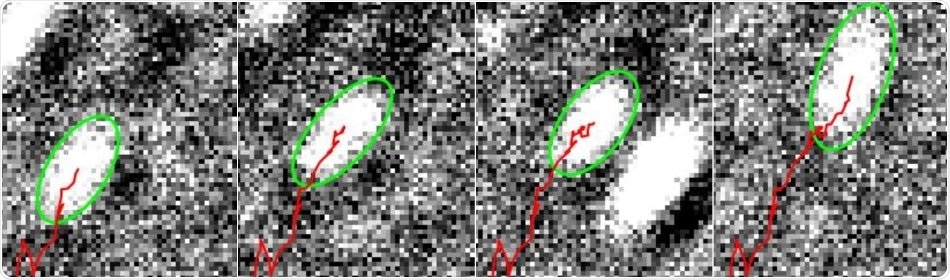
Myosin being tracked by the software. Image Credit: University of Warwick
In a study recently published in the Biophysical Journal, researchers from the University of Warwick’s Department of Physics and Warwick Medical School have utilized a novel microscopy method to examine the molecular motors that exist within cells.
These molecular motors enable the cells to move and reshape on their own, possibly offering a new insight that could lead to the development of innovative smart materials.
Myosin is a type of protein that creates the motor filaments that, in turn, provide stability to cells and play a role in remodeling the actin cortex within the cell.
The actin cortex is relatively similar to the cell’s backbone and gives shape to the cell, whereas the myosin filaments are analogous to muscles. By “flexing,” the myosin filaments allow the cell to apply forces outside of it and thus spread.
When the cell wants to contract or exert stresses on its neighbouring cells or tissue it will form stress fibres using an array of myosin and actin filaments to perform an action similar to our own muscles. In fact, our muscles are made up of the same molecules.”
Dr Darius Köster, Study Corresponding Author, Warwick Medical School, University of Warwick
Dr. Köster continued, “But they can also form other structures. A cell is constantly remodeling itself and all these proteins are undergoing a turnover because they have a limited lifetime. A number of processes require the local remodeling of the cortex, for example, if the cell wants to move around it uses actin remodeling and myosin.”
A method, known as optical fluorescence microscopy, is often used by biologists to study biological molecules. In this method, a fluorescent protein is added to the molecule, which is subsequently activated so that it emits light that can be identified and examined. However, this method has one disadvantage—it causes phototoxicity and thus destroys the molecule while one looks at it.
To overcome this problem, the researchers from the University of Warwick teamed up with the Kukura laboratory at the University of Oxford to examine a minimal cell cortex with the help of a method known as interferometric scattering microscopy, or iSCAT for short. The researchers do not need to add fluorescent proteins to the molecules in this technique, which utilizes scattered light from the source molecule.
By quantifying the interference between that light and the light emitted by the glass surface on which the sample was placed on, the researchers can create more scalable and sensitive imagery while restricting the amount of light that the molecule is subjected to.
The researchers can observe objects at the optical resolution limit, that is, at 0.0002 mm or 200 nm, while the muscle myosin filaments usually measure 0.001 mm long.
Our minimal cell cortex rearranges itself over periods of tens of minutes, so you have to be able to follow activity at short timescales, but over long periods. And that’s difficult using conventional fluorescent microscopy.”
Dr Darius Köster, Study Corresponding Author, Warwick Medical School, University of Warwick
Dr Köster added, “We wanted to find out whether myosin aligns to actin and exerts tensile forces, or if it can cross over to several filaments and bring them together to contract them. With this new technique what we actually see is different regimes of fluctuations in the myosin filaments. Myosin has a dumbbell structure and when only one side binds to actin it fluctuates a lot. If both sides bind down then they are much stiffer and don’t fluctuate as much.”
“We found that myosin can behave in both regimes, something that wasn’t possible to observe before. There are very few studies of these ensembles, others have studied single molecules at this timescale but you could only theorize how a motor ensemble works,” Dr. Köster further added.
Lewis Mosby is a Ph.D. student in the Department of Physics and Warwick Medical School jointly supervised by Dr. Marco Polin and Dr. Anne Straube. At present, Mosby is exploring intracellular cargo transport with the help of a grant from the Leverhulme Trust managed by Dr. Straube.
Lewis Mosby is the lead author of the study and he programmed a novel particle tracking software to detect the orientation as well as the position of the myosin filaments.
We used a code that was developed to detect galaxies and other objects in the sky, so it was fascinating to apply that to the exact opposite end of the scale.”
Lewis Mosby, Study Lead Author, Department of Physics, Warwick Medical School, University of Warwick
In addition to offering a better understanding of the overall function of the cortex of mammalian cells, the molecular motors serve as a model for innovative new technologies.
“This is a nice example of a self-organizing composite with a range of different dynamics but all built-in due to the properties of the proteins. That’s an interesting feature if you are looking at, for example, smart materials. It’s not only self-organizing but also active, taking on different shapes depending on the consumption of energy,” concluded Dr. Köster.
Source:
Journal reference:
Mosby, L. S., et al. (2020) Myosin II Filament Dynamics in Actin Networks Revealed with Interferometric Scattering Microscopy. Biophysical Journal. doi.org/10.1016/j.bpj.2020.02.025.