Specific sequences of DNA are typically cut using CRISPR-Cas9 that has become one of the most effective and convenient biotechnology tools today.
Beginning from Streptococcus pyogenes Cas9 (SpCas9), a host of variants have been designed and used in experiments across the world.
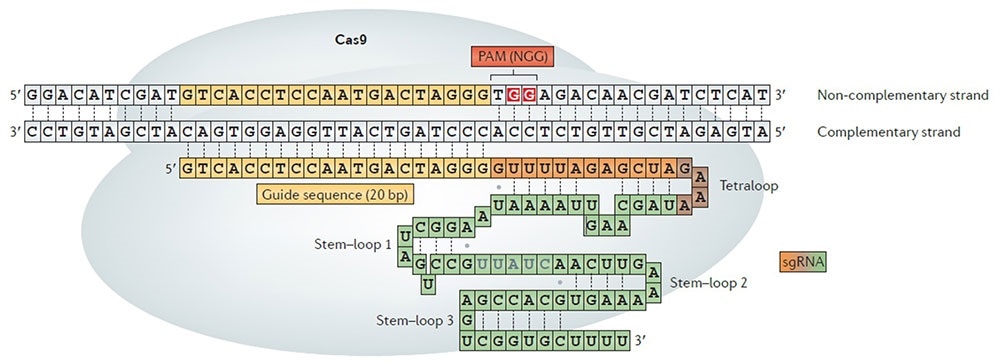
Schematics of the CRISPR-Cas9 system. A guide RNA guides Cas9 to the target DNA sequence, which is followed by the short protospacer adjacent motif (PAM). Researchers across the globe have been adopting this technology to cut DNA at desired positions Image Credit: Kim, H., & Kim, J. S. Nature Reviews Genetics, 2014.
While all these systems are currently used to target and cleave a particular sequence of DNA, they also show comparatively high off-target activities that have possibly harmful effects.
The team of researchers, from the Center for Nanomedicine, within the Institute for Basic Science (IBS, South Korea) and headed by Professor Hyongbum Henry Kim, has realized the most widespread high-throughput analysis of CRISPR-Cas9 activities.
The researchers designed deep-learning-based computational models that estimate the activities of the SpCas9 variants for different sequences of DNA. The study, published in the Nature Biotechnology journal, serves as a useful guide for choosing the most suitable variant of SpCas9.
The study exceeded all the earlier reports that had assessed only a total of three Cas9 systems. After comparing 13 variants of SpCas9, the IBS scientists defined which four-nucleotide sequences can be employed as a protospacer adjacent motif (PAM). PAM is a short DNA sequence needed for Cas9 to cut and is placed instantly after the cleavage-targeted DNA sequence.
In addition, the scientists assessed the specificity of six different, high-fidelity variants of SpCas9 and observed that evoCas9 exhibits the highest specificity, whereas the original wild-type SpCas9 exhibits the lowest specificity.
Even though evoCas9 is highly specific, it still shows minimal activity at several target sequences: Such results suggest that other high-fidelity Cas9 variants could be favored based on the DNA target sequence.
On the basis of these outcomes, the IBS team created a computational tool, called DeepSpCas9variants, to estimate the activities of the SpCas9 variants. When users access this public website, they may enter the required DNA target sequence, identify the most appropriate SpCas9 variant, and fully leverage the CRISPR technology.
We began this research when we noticed the critical lack of a systematic comparison among the different SpCas9 variants. Now, using DeepSpCas9variants, researchers can select the most appropriate SpCas9 variants for their own research purposes.”
Hyongbum Henry Kim, Professor, Center for Nanomedicine, Institute for Basic Science
Source:
Journal reference:
Kim, N., et al. (2020) Prediction of the sequence-specific cleavage activity of Cas9 variants. Nature Biotechnology. doi.org/10.1038/s41587-020-0537-9.