The genetic scissors, known as CRISPR/Cas9, the finding that received the 2020 Nobel Prize in Chemistry, occasionally cut in places that they are not intended to target.
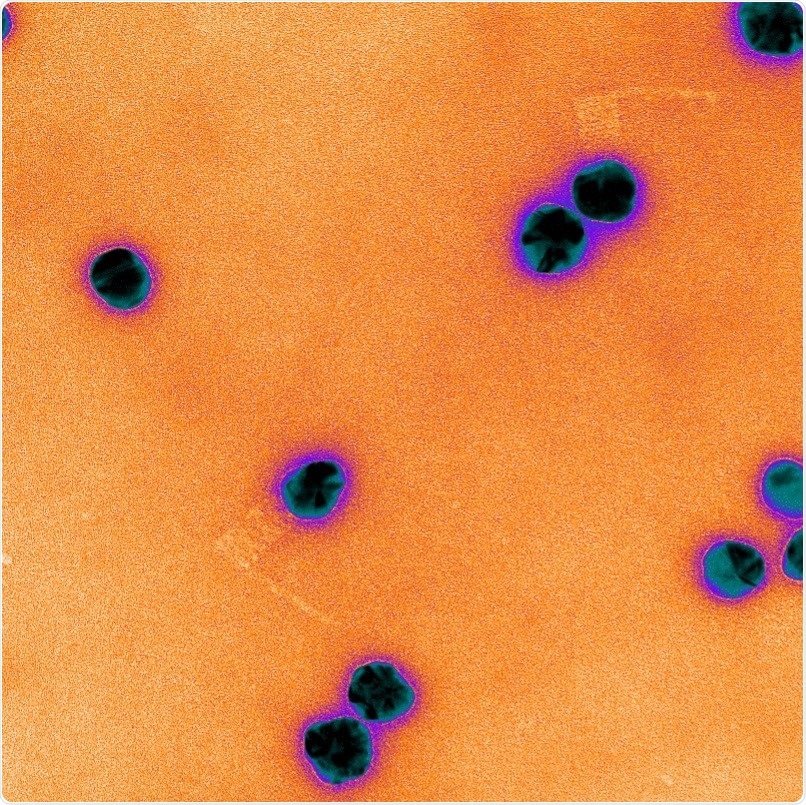
Electron microscopy image of DNA origami rotor arms, which are the faint orange “L’s” attached to the purple tracking particles. Image Credit: Julene Madariaga Marcos.
While CRISPR has fully altered the pace of fundamental research by enabling researchers to rapidly edit genetic sequences, this technique works so quickly that it is difficult for researchers to observe what sometimes goes wrong and find out how to enhance it.
Julene Madariaga Marcos, a Humboldt postdoctoral fellow, and collaborators in the laboratory of Professor Ralf Seidel at Leipzig University in Germany discovered a method to examine the ultra-fast movements of CRISPR enzymes, which will allow scientists to interpret how they identify their target sequences in an attempt to enhance the specificity.
Madariaga Marcos will present the study at the 65th Annual Meeting of the Biophysical Society. on February 23rd, 2021.
To utilize CRISPR enzymes to edit gene sequences, researchers can customize them to target a particular sequence within the three billion DNA base pairs found in the human genome. At the time of target recognition, CRISPR enzymes untwist the strands of DNA to identify a sequence that is complementary to the attached RNA sequence of CRISPR.
However, the RNA occasionally matches the sequences of DNA that are not quite complementary. Hence, to overcome this unintentional match, researchers should be able to see how CRISPR is working along with separate DNA base pairs; however, this process is too quick and hard to visualize.
To quantify CRISPR's activities on an ultrafast timescale, Madariaga Marcos and collaborators turned to DNA origami, which employs special sequences of DNA to create complex 3D nanostructures rather than a simple double helix.
DNA origami can be used in nanoelectronics, drug delivery, and even art. With the help of DNA origami, the team constructed rotor arms out of DNA so that through a high-speed camera on a microscope, they could observe the untwisting of the DNA by CRISPR enzymes, making the rotor arm rotate like helicopter blades.
Through this system, the researchers were able to quantify the different responses to matches and mismatches inside the DNA sequence.
We are able to directly measure the energy landscape of CRISPR/Cascade when it interacts with DNA for the first time.”
Julene Madariaga Marcos, Humboldt Postdoctoral Fellow, Leipzig University
This method will allow investigators to interpret CRISPR enzymes, and how they eventually land on their match. In this way, they can find out how to improve CRISPR so that it makes fewer matches that are off-target. In the days to come, Madariaga Marcos is keen on “developing more tools and methods for studying these gene-editing processes in new ways and at a more detailed level.”