What provoked your research into the impact of diseases on the immune system?
As a child, I used to hear my mom talking about T cells, how these cells would defend us against microbes, and how important these “soldier cells” were to us. It was due to a natural bias that after graduating in Mathematics at the University of Buenos Aires, Argentina, I came to France to study genetic epidemiology in one of the most prestigious laboratories studying human predisposition to infectious diseases.
But regardless of my academic past, I think many of us, including myself, are interested in understanding the main questions that have intrigued humans; Who are we? Where are we from? Where do we go? And these questions, although extremely broad, are related to human evolution.
When finishing my Ph.D. I, therefore, moved to one of the best laboratories in the field of human genetic evolution, my current host lab, to study susceptibility to infectious diseases from an evolutionary perspective.
The laboratory I work in is interested in questions related to human adaptation across time and space, and among the main past events that have forced humans to adapt are the ones related to infectious diseases, that have accompanied us since the very beginning of our species.
Following this approach, I focused on gene variants that might have contributed to the rise and fall of past epidemics.

Genetic Evolution. Image Credit: PopTika/Shutterstock.com
How have diseases and infections evolved?
This is one of the main questions that we, and many other groups, try to answer, or at least bring some light to it. As I was saying before, co-evolution with pathogens is key in explaining human movements, expansions, adaptation to new environments, and the success of the species. Old microbes have reached some levels of co-adaptation with us (e.g. microbiota) as we co-exist in relatively perfect harmony. This is the ultimate goal of microparasites: perfect balance to preserve survival.
However, pathogens evolve as humans evolve, and new pathogens emerge, either because we move to new environments, modify the environment, or simply by chance. Reconstruction of ancient pathogens is also an emerging discipline and we will understand more of this in the near future.
What is tuberculosis and how can this cause infection?
Tuberculosis is an infectious disease caused by mycobacteria known as Mycobacterium tuberculosis and infects humans as well as other species. Contact with the mycobacteria can lead to infection and ultimately disease.
Animals can be a reservoir for the mycobacteria in the same way humans can be a reservoir for the mycobacteria. The evolution of the mycobacteria is important to study as we might learn how it appeared, co-evolved, and adapted to us.
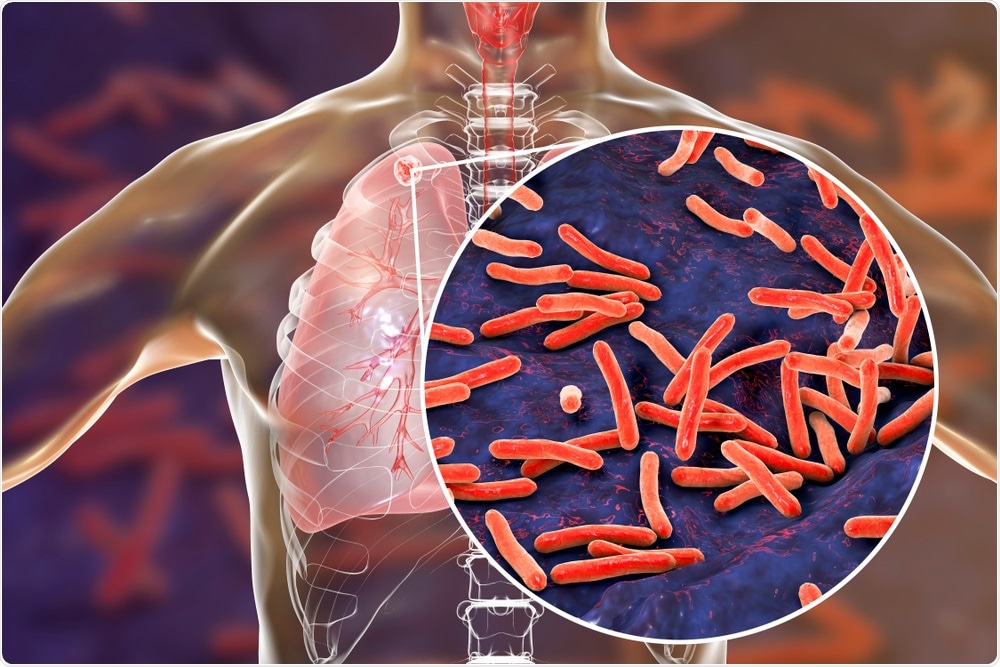
Tuberculosis. Image Credit: Kateryna Kon/Shutterstock.com
Can you describe how you carried out your latest research into tuberculosis and the immune system?
Our logic is that pathogens have always “pressured” humans. Virulent pathogens have been the cause of the death of massive portions of past populations. As the head of my lab says, we are here because our ancestors have survived some of these past epidemics, black death, Spanish flu, Salmonella in Mesoamerica, etc., and their genes, as opposed to the genes of those who died, might have carried, in average, an advantage.
If a variant strongly increases in frequency in a population over time it might be because it was beneficial to survival, for example, it might protect against an infectious agent. In our case, we had this variant in the TYK2 gene that we found, during my Ph.D., to be a predisposition to tuberculosis.
We found that by looking at modern databases of tuberculosis cases and by proper immunological testing in the laboratory. If this variant is so deleterious in the context of tuberculosis, and because tuberculosis was endemic only 150 years ago in Europe (and even more recently), we wondered if there was a signature in the evolution of this variant that could inform us about past tuberculosis epidemics.
To test for this, we performed extensive computer simulations under many different scenarios of evolution to test which one could have been the more realistic according to what we know happened for this gene variant.
We used ancient human DNA, which is the DNA of humans that have lived at different epochs of the past 10,000 years ago or so, and we evaluated the variations in frequency for this gene variant over time. The simulations that better matched the distributions of frequency obtained from these ancient genomes were the ones that better accounted for what happened at the evolutionary level.
What did you discover?
What we found is that the variant dramatically decreased in frequency only after the Bronze Age in Europe (this could only be explained due to some external pressure, like that of the mycobacterium tuberculosis). Homozygotes for this gene variant were negatively selective, which means that those carrying 2 copies of this variant died faster (before reaching the reproductive age) as compared with those having 1 or 0 copies of this variant.
We also discovered that we could date the start of this negative selection. Individuals with 2 copies of this gene variant started to die faster than the rest around 2,000 years ago (despite the gene variant existing for 30,000 years), which suggests that strong tuberculosis epidemics have started around this period. If the mycobacteria existed before it did not have such a strong effect on Europeans compared to its effect over the last 2,000 years.
See fullscreen
How has your research helped in the field of evolutionary genetics?
This research is a sort of proof-of-concept that we can delineate variants with a strong impact on modern health by studying its evolutionary history with ancient DNA.
Do you believe that with more research, we can further understand how genetics can influence the immune system?
Yes, of course, and this is something that my so far two host labs have been working on for many years. What predisposes us to infectious diseases?
For sure human genetics has a strong role in it and though hints are already on the table, there is much still to be discovered.
How could your research be used to help further understand the current COVID-19 pandemic?
The current COVID-19 pandemic is quite peculiar compared to past pandemics since it has had a major impact on the elderly but not so much on the younger populations. Since the elderly will not largely contribute, genetically speaking, to the next generations, we do not expect this pandemic to have an effect that could be traced back to the gene level.
However, it is interesting to see that some variants that were selected in the past by our species are beneficial today to fight against COVID. I refer here to the example of a Neanderthal haplotype that protects us against severe COVID disease. If we kept this variant, inherited from Neanderthal, in our genomes it is likely because it was already beneficial in the past.
What are the next steps in your research?
The next step here is to assess other variants that have also decreased or increased strongly in frequency and learn about them in the context of past events, such as epidemics. We might find other extinct variants that predispose or protect us against infectious agents.
The ultimate goal here is to provide the tools for the development of therapeutic treatments.
Where can readers find more information?
Map of burial places from where ancient DNA has been extracted so far: http://umap.openstreetmap.fr/en/map/ancient-human-dna_41837#4/14.77/9.45
Article on our paper by science journalist: https://www.sciencemag.org/news/2021/03/how-tuberculosis-reshaped-our-immune-systems
About Dr. Gaspard Kerner
Gaspard is Argentinian and French and is currently working as a post-doc in Lluis Quintana-Murci’s laboratory, Human Evolutionary Genetics at Institut Pasteur, Paris. He completed a Ph.D. in human predisposition to tuberculosis in Institut Imagine, Paris under the supervision of Laurent Abel in the lab co-directed by Laurent Abel and Jean-Laurent Casanova, where he was able to understand the role of the TYK2 gene variant in its susceptibility to humans forms of tuberculosis.
Dr. Kerner has a mathematical background from his years in Argentina and uses his diverse background (math, population genetics, genetic epidemiology) to develop new methods to better understand our co-evolution with pathogens.