A single-celled alga removes non-essential pieces by undergoing genome surgery. This may result in the most effective cellular factory for creating sustainable and renewable biofuels from carbon dioxide and sunlight.
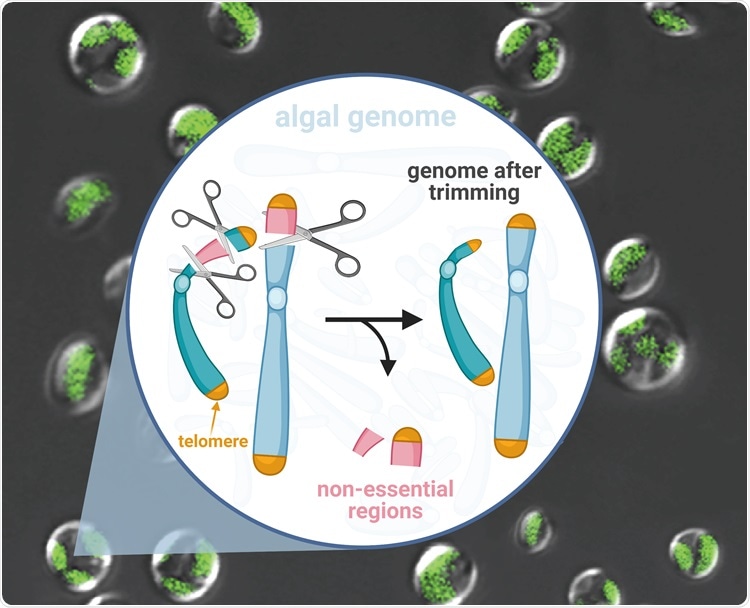
Hundred-kilobase fragment deletions in microalgae by Cas9 cleavages. This figure was made using BioRender. Image Credit: Yang Liu.
A research team from the Qingdao Institute of BioEnergy and Bioprocess Technology (QIBEBT) of the Chinese Academy of Sciences has extracted a hundred-kilobase genome from a kind of oil-producing microalgae, knocking out genes that are not crucial for it to function. By doing so, the team has developed a “genome scalpel” that is capable of trimming microalgal genomes quickly and creatively.
The resultant “minimal genome” microalgae could be used as a model organism for additional research into the biological and molecular role of each gene, or as a “chassis” strain for synthetic biologists to boost the tailored production of biomolecules, like bioplastics or biofuels. The study results were published in The Plant Journal.
The development of a “minimal genome,” or a genome devoid of all duplicated or seemingly non-functional “junk genes,” can be extremely useful for exploring the underlying questions about the genetic function and for engineering cell factories that generate useful compounds.
These minimal genomes have been produced for simple organisms, but seldom for eukaryotic organisms, such as plants or algae. In the case of higher eukaryotes, “junk” regions can occupy up to 70% of the genome. As a matter of fact, deleting what seems to be simply “junk genes” can have a detrimental impact on the organism or even destroy it.
Now, for the first time, the QIBEBT team has created a genome with targeted deletions, each measuring a hundred kilobase in size, for a kind of algae species called Nannochloropsis oceanica.
N. oceanica are essentially single-celled algae (microalgae) that have immense potential for producing biomaterials, biofuels, and other platform chemicals in a sustainable and renewable manner while decreasing the emissions of greenhouse gas. But to realize the potential of such microalgae, elaborate genetic engineering of the organism is required to minimize production costs and maximize yields.
The QIBEBT research team initially detected the non-essential chromosomal areas—those in which genes were seldom activated or expressed. They discovered 10 such “low-expression regions” or LERs and then applied the CRISPR-Cas9 gene-editing method to remove two of the largest LERs, measuring more than 200 kilobases in size.
“Despite the all snipping, the microalgae still showed essentially normal growth, lipid content, fatty acid saturation levels, and photosynthesis. In some cases, there was even a slightly higher growth rate and biomass productivity than the organism in the wild,” stated the study first-author Qintao Wang from the Single-Cell Center (SCC) in the QIBEBT.
We interestingly found normal telomeres in the telomere-deletion mutants of Chromosome 30. This phenomenon implies the loss of distal part of chromosome may induce telomere regeneration.”
Jian Xu, Study Corresponding Author, Single-Cell Center, Qingdao Institute of BioEnergy and Bioprocess Technology, Chinese Academy of Sciences
The significantly snipped genome in Nannochloropsis can already function as a closer-to-minimal genome, which can act as the chassis strain for tailored biomolecule production using more metabolic engineering atop this chassis.
Now after demonstrating that they can strip down the genome of such a complex eukaryote, the team wants to find out if they can further snip out the LERs and other non-lethal areas to design a truly minimal Nannochloropsis that makes biofuels from carbon dioxide with maximum efficiency.
Source:
Journal reference:
Wang, Q., et al. (2021) Genome engineering of Nannochloropsis with hundred‐kilobase fragment deletions by Cas9 cleavages. The Plant Journal. doi.org/10.1111/tpj.15227.