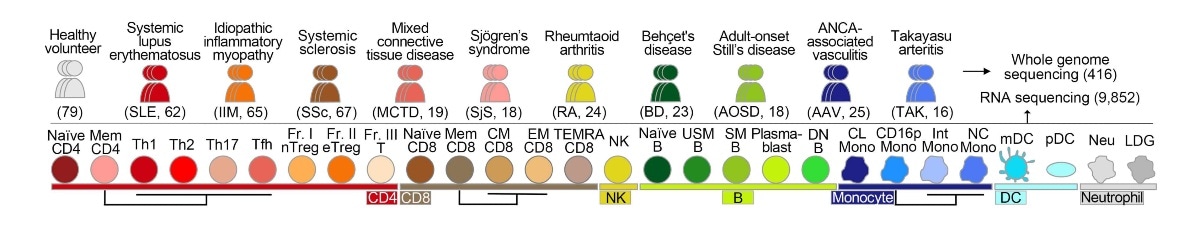
Full genome sequencing was completed for a total of 79 healthy volunteers and 337 patients diagnosed with one of 10 different immune-mediated diseases (top row) to help build a new genetic atlas, ImmuNexUT. Researchers isolated 28 different immune cell types from their blood samples (bottom row) and measured the expression of genes inside each cell type. Image Credit: Mineto Ota.
Researchers also expect that this atlas of immune-related genome data can be useful in the study of infectious diseases such as COVID-19.
Ota is the study lead author and Project Research Associate, and the study was published recently in Cell. The project was conducted in collaboration with researchers from RIKEN research institution and Chugai Pharmaceutical Co., Ltd.
Several previous studies contrasted the complete genome sequences of patients with disease diagnoses to those of healthy individuals. All DNA sequence variants discovered in these genome-wide interaction studies are regarded as “associated” with the disorder.
Many of the variants discovered in association studies are not found in genes, the fundamental units of heredity, but instead in regions of DNA that control the “off” or “on” expression of genes. The majority of the human genome is made up of regulatory DNA rather than genes.
Experts may be aware that a portion of DNA is involved in gene control, but they may not understand what or how it does, or even which genes it controls.
A different kind of genome study known as expression quantitative trait loci (eQTL) research aims to link variations in DNA sequence to differences in gene expression to reveal the role of regulatory DNA.
Using eQTL information, researchers may make more informed assumptions about the function of regulatory DNA sequences, how variants in the regulatory sequence can influence the expression of the genes it controls, and how variations in gene expression cause disease.
Although other immune-focused eQTL studies have been conducted, previous research attempts involved only healthy volunteers and looked at a small number of cell types.
Inflammatory conditions create very different physical characteristics in immune cells compared to the same cells in a healthy condition. The genetic variants associated with immune conditions may only function in the diseased state, so for this type of genetic study, it was very important to get samples from real patients.”
Mineto Ota, MD, PhD, Clinical Rheumatologist and Expert in Functional Genomics, University of Tokyo
The researchers sequenced the whole genomes of 79 healthy volunteers and 337 patients with immune-mediated disorders from 10 different groups, including systemic lupus erythematosus, systemic sclerosis, and rheumatoid arthritis. All the volunteers were of Japanese ancestry.
Comprehending immune-mediated diseases is difficult. This is because though each condition is clinically unique, there are several overlaps, and patients with the same diagnosis may respond with highly varied symptoms.
Due to all this diversity, there’s a limit to how much you can learn studying one immune-mediated disease at a time. But if we study 10 diseases together, it gives a bigger picture about these types of diseases.”
Mineto Ota, MD, PhD, Clinical Rheumatologist and Expert in Functional Genomics, University of Tokyo
Researchers isolated 28 different types of immune-related cells from blood samples of volunteers and analyzed gene expression in those cells after completing full genome sequencing.
The Immune Cell Gene Expression Atlas from the University of Tokyo (ImmuNexUT) was the name given to the database developed as a result of this study.
“We see that each immune cell type has distinctive eQTL results in the atlas, which can tell us how gene regulation differs between cell types and exactly which cell type is important for developing which disease,” said Ota.
ImmuNexUT has grown to become the largest eQTL data collection created by volunteers of East Asian ancestry.
“In this field so far, large-scale genomic and functional genomic studies have been mainly conducted with European donors, although population diversity is critical for precise understanding of the genomic functions. This eQTL atlas of Japanese subjects is also meaningful for overcoming this European-centric bias and further investigating the functions of DNA variants in combination with European data sets,” added Ota.
The team expects that the ImmuNexUT database will continue to expand and potentially lead to improved patient outcomes.
Source:
Journal reference:
Ota, M., et al. (2021) Dynamic landscape of immune cell-specific gene regulation in immune-mediated diseases. Cell. doi.org/10.1016/j.cell.2021.03.056.