Researchers from the European Molecular Biology Laboratory (EMBL) and the German Cancer Research Center (DKFZ) demonstrated a new technique to create metabolic profiles of individual cells.
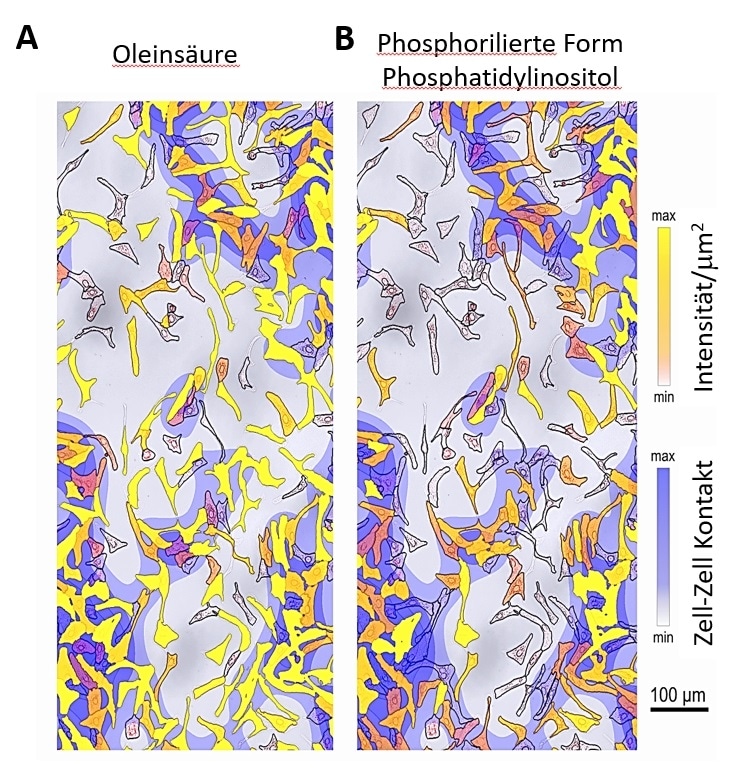
Image Credit: © DKFZ/Heikenwälder.
The newly created technique combines a specific form of mass spectroscopy and fluorescence microscopy to analyze over a thousand individual cells and more than a hundred lipids and metabolites in an hour. The scientists anticipate that this newly devised technique can provide solutions to different biomedical questions.
Biomedical disciplines currently focus on individual cell metabolites, which were once considered as degradation products or building blocks for the synthesis of complex cellular molecules. The role of these metabolites in providing support and determining central cell functions as signaling molecules, thereby making a significant contribution in maintaining the healthy balance of the body, has been currently understood.
Metabolites modulate epigenetics, they regulate the immune system, control inflammation, and are thus also involved in carcinogenesis. Therefore, the analysis of metabolite profiles has become the focus of interest in many research disciplines during the last few years.”
Mathias Heikenwälder, German Cancer Research Center
Individual cells of tissues or organs do not have a uniform metabolite profile; however, they often exhibit pronounced heterogeneity in the broad profile of different metabolites and lipids. The individual metabolite profile of a cell is also based on its localization within the organ.
Organ or tissue conditions can be assessed by evaluating the metabolic profile of a large number of individual cells and parallelly documenting their localization within the tissue network. The “SpaceM” developed by the Heidelberg researchers provides a novel method to perform these analyses with high throughput for a wide range of adherent cells, specifically, liver cells.
The innovative technology is based on a combination of fluorescence microscopic images and a unique form of mass spectrometry known as MALDI-imaging (Matrix Assisted Laser Desorption/Ionization). The use of this innovative technique enabled the Heidelberg scientists to detect over 1000 individual cells and more than a hundred metabolites in an hour. Subcellular scale measurements are also possible in the future.
To substantiate the newly devised technique, the researchers investigated human liver cells stimulated with fatty acids and proinflammatory cytokines as an in vitro model of widespread liver diseases—the Non-Alcoholic Fatty Liver Disease and Non-Alcoholic Steatohepatitis.
The researchers used SpaceM to obtain metabolite profiles of nearly 30,000 individual cells. Steatosis, an inflammatory change, was observed in a quarter of cells indicated by a uniquely modified lipid profile. Further stimulation of the cells with the pro-inflammatory messenger molecule IL17A made the entire cells switch to the inflammatory phenotype.
The researchers confirmed the same sequence of changes in the development of inflammatory liver disease in mice through standard measurement methods.
SpaceM offers novel capacities for spatial single-cell metabolomics. Yet is cost-efficient and uses existing instrumentation. We anticipate that the method and open-source algorithms will democratize single-cell metabolomics worldwide to answer a variety of pressing biomedical questions.”
Theodore Alexandrov, European Molecular Biology Laboratory
Source:
Journal reference:
Rappez, L., et al. (2021) SpaceM reveals metabolic states of single cells. Nature Methods. doi.org/10.1038/s41592-021-01198-0.