Cell division is a vital mechanism underlying biological growth and repair. Cell biologists trace this mechanism by examining chromosomes—structures made of DNA containing the genetic material of an organism.
Application of deep learning artificial intelligence (AI) for image classification requires a high level of expertise in machine learning. Now, researchers from Japan have created an AI image classifier that can be customized and readily used by non-experts. Image Credit: Kiyotaka Nagaki from Okayama University.
Progress in the field of microscopy along with automation has enabled scientists to get better images of chromosomes in a short duration. But most of the examination is still performed manually and is mostly a tedious task. This is particularly true for plants that show a huge variety in chromosome sizes and numbers.
Scientists from Japan took a different approach. The study headed by associate professor Kiyotaka Nagaki from Okayama University, Japan, employed deep learning artificial intelligence (AI) to classify chromosomal images from various plant species. The exciting part is that the researchers have demonstrated that even non-experts can use AI effortlessly. The research was published in the journal Chromosome Research.
Dr. Nagaki elaborates on how it was achieved.
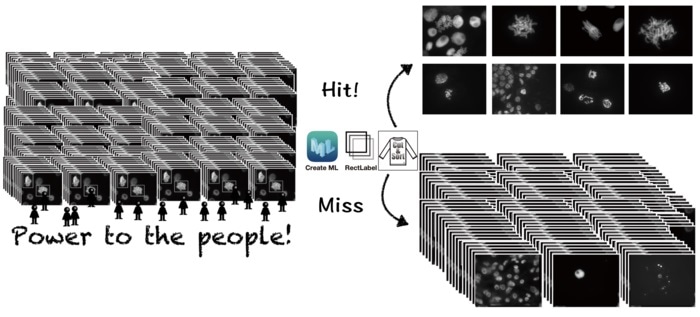
Classifying images using AI usually requires a high level of computer knowledge. What we did is build AI models on a McIntosh computer with the CreateML app suitable for our own image samples. Moreover, the AI can be trained to become an order-made image classifier for any variety of images that suits one’s purpose.”
Kiyotaka Nagaki, Associate Professor, Okayama University
The researchers utilized chromosomal images to train the deep learning models to identify images or parts of images where cells undergo “mitosis”—a mechanism where a single cell divides into two identical daughter cells. The researchers evaluated its detection accuracy with test images depending on the number of cells the model correctly classified.
The scientists later tested the models with images having mitotic cells from plant species that were not involved at the time of the training. They observed that the models precisely differentiated mitotic cells in the images. Additionally, the method also functioned effectively for cells in tissue sections and a diverse mechanism of cell division.
The observations show that the deep learning pipeline created by the researchers can be effortlessly and accurately employed by non-data researchers across various disciplines, highly simplifying and boosting up the task of image analysis.
Moreover, the scope of the current method can be prolonged towards much complex examination like the recognition of chromosome aberrations and the formation of novel advisory systems for object identification and classification.
There are more trivial classifications in our lives than one might imagine. Automating such classifications by entrusting them to an AI can not only eliminate fluctuations caused by individual differences but also save many valuable research hours. Streamlining such trivial classifications make extensive image-based studies more reproducible and reliable.”
Kiyotaka Nagaki, Associate Professor, Okayama University
Undoubtedly as Dr. Nagaki says, “a deep learning sorter that anyone can use,” can be the solution to comprehend different nuances of biological species.
Source:
Journal reference:
Nagaki, K., et al. (2021) Effectiveness of Create ML in microscopy image classifications: a simple and inexpensive deep learning pipeline for non-data scientists. Chromosome Research. doi.org/10.1007/s10577-021-09676-z.