Cryogenic electron microscopy (cryo-EM) allows scientists to investigate the assembly of DNA replication machinery where DNA is damaged.
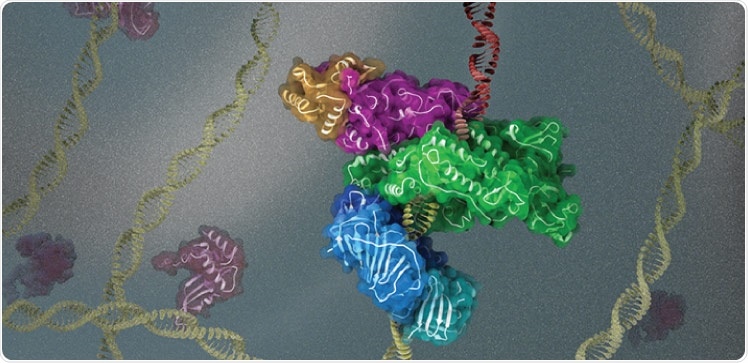
KAUST scientists use cryogenic electron microscopy to investigate the 3D structure and function of key protein complexes involved in DNA replication and repair. Image Credit: © 2021 King Abdullah University of Science & Technology; Heno Hwang.
Cellular DNA is constantly exposed to both exogenous and endogenous DNA-damaging agents like UV radiation and reactive oxygen species. To decrease the biological effects of DNA damage, living organisms have evolved processes to endure and repair DNA damage to assure that genetic information is precisely inherited.
One of those mechanisms, known as translesion synthesis (TLS), permits DNA replication to advance through unrepaired DNA lesions.
TLS includes highly precise DNA synthesizing enzymes (replicative DNA polymerases) temporarily replaced with specific, low-fidelity TLS polymerases that ensure cell survival by introducing mutations. The translesion synthetic and mutagenic activity of TLS polymerases might cause the normal cells to become cancerous, or cancer cells become drug-resistant.
The Y-family TLS polymerase Pol κ is capable of carrying out DNA synthesis across numerous damaged bases and is employed to DNA lesions by proliferating cell nuclear antigen (PCNA). Earlier researches indicate that PCNA is controlled by ubiquitination.
The addition of a single ubiquitin molecule at lysine residue 164 (K164) of PCNA facilitates the recruitment and retention of TLS polymerases to damage sites, but the structural basis of the interaction between these polymerases and ubiquitinated PCNA is poorly understood.”
Alfredo De Biasio, Structural Biologist, King Abdullah University of Science & Technology
Since 2018, De Biasio’s team associated with the laboratory headed by Samir Hamdan, a specialist in the single-molecule analysis of human DNA replication. The researchers were utilizing cryo-EM to analyze the three-dimensional structure and function of major protein complexes engaged in DNA replication and repair.
The recent research elaborates cryo-EM reconstructions of full-length human Pol κ attached to DNA, an incoming nucleotide, and mono-ubiquitylated PCNA or unmodified PCNA at near-atomic resolution. The researchers identified that when DNA is absent, the structure of Pol κ attached to PCNA is greatly flexible, indicating that linkage to DNA is needed to form a rigid and active complex.
A senior researcher in Hamdan’s group and the co-lead author of the research, Muhammad Tehseen, carried out major functional researches illuminating how PCNA ubiquitination regulates the activity of Pol κ.
Our data provide a structural framework to explain how PCNA recruits a Y-family TLS polymerase to sites of DNA damage.”
Muhammad Tehseen, Study Co-Lead Author and Senior Researcher, King Abdullah University of Science & Technology
Due to the high degree of domain conservation between Y-family polymerases, certain structural features seen in the Pol κ-DNA-PCNA complex might be applicable to other TLS polymerase complexes.
By understanding the interactions between the proteins forming these complexes and how they are regulated, we can identify ways to reduce or increase their function for medical applications.”
Muhammad Tehseen, Study Co-Lead Author and Senior Researcher, King Abdullah University of Science & Technology
Source:
Journal reference:
Lancey, C., et al. (2021) Cryo-EM structure of human Pol κ bound to DNA and mono-ubiquitylated PCNA. Nature Communications. doi.org/10.1038/s41467-021-26251-6.