Artificially synthesizing genomes has several applications, including medical research and industrial strains.
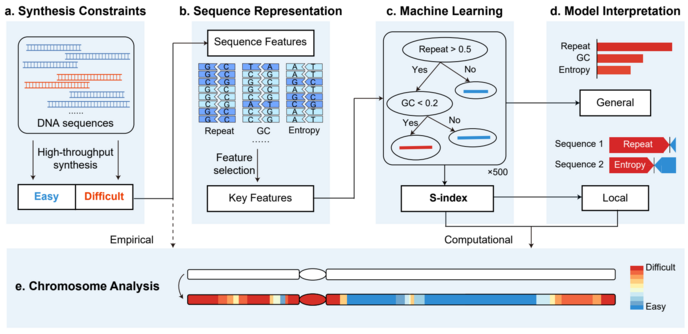 A, Collection of the DNA sequences obtained from high-throughput synthesis. The sequences were classified into easy-to-synthesize (blue) or difficult-to-synthesize (red). B, Graphical representations of DNA sequences: repeat, GC content, information entropy, and other types of features. Key features were identified from these sequence features by machine learning methods. C, The XGBoost algorithm utilized to build the classification model and calculate the S-index. D, Methods used to interpret the model. The feature contributions were quantified according to the global importance scores and local SHAP explanations. e, Application of the S-index on a specific chromosome. The heatmap indicates the synthesis difficulties for the different fragments, which range from difficult (red) to easy (blue). The white sequences indicate the unanalyzed chromosome sequence. Image Credit: Science China Press
A, Collection of the DNA sequences obtained from high-throughput synthesis. The sequences were classified into easy-to-synthesize (blue) or difficult-to-synthesize (red). B, Graphical representations of DNA sequences: repeat, GC content, information entropy, and other types of features. Key features were identified from these sequence features by machine learning methods. C, The XGBoost algorithm utilized to build the classification model and calculate the S-index. D, Methods used to interpret the model. The feature contributions were quantified according to the global importance scores and local SHAP explanations. e, Application of the S-index on a specific chromosome. The heatmap indicates the synthesis difficulties for the different fragments, which range from difficult (red) to easy (blue). The white sequences indicate the unanalyzed chromosome sequence. Image Credit: Science China Press
Researchers are continually making advances in the depth and breadth of genome design and synthesis, from Craig Venter’s team’s synthesis of the artificial life JCVI-syn1.0 in 2010 to the rewriting and synthesis of the prokaryotic E. coli genome, and to the Sc2.0 project’s artificial synthesis of the yeast genome.
A limitation of the use and promotion of artificial genome synthesis technology is the constant challenge of synthesizing specific gene segments, which eventually prevents artificial chromosomes from being completed.
To solve this problem, the Tianjin University team led by Professor Yingjin Yuan has created an interpretable machine-learning framework that can foresee and measure the complexity of chromosome synthesis and offer recommendations for improving chromosome design and synthesis procedures.
By analyzing data from a vast number of known chromosome segments, the study team developed an effective feature selection approach and found six important sequence characteristics that encompass energy and structural information during DNA chemical production and assembly.
Using these findings as a foundation, the researchers created the eXtreme Gradient Boosting (XGBoost) model, which can accurately anticipate the synthesis challenges of chromosome fragments.
The model’s high accuracy and prognostic ability were demonstrated by its AUC (area under the receiver operating characteristic curves), which was 0.895 in cross-validation and 0.885 on an independent test set in association with a DNA synthesis company.
To assess and explain the synthesis difficulties of chromosomes, the study team created the Synthesis Difficulty Index (S-index), which is based on the SHAP method.
The study discovered that the synthesis difficulties of various chromosomes differed significantly, and the S-index could quantitatively articulate the causes of synthesis difficulties for some gene fragments.
This information provided a foundation for chromosome sequence design and synthesis and increased the effectiveness and success rate of designer chromosome synthesis.
This accomplishment gives chromosomal engineering and genome rewriting researchers a useful tool, and it is anticipated to offer more comprehensive instructions and assistance for chromosome design and synthesis.
Source:
Journal reference:
Zheng, Y., et al. (2023). Machine learning-aided scoring of synthesis difficulties for designer chromosomes. Science China Life Sciences. doi.org/10.1007/s11427-023-2306-x