A new genome editing technology was developed by GAO Caixia’s group from the Institute of Genetics and Developmental Biology of the Chinese Academy of Sciences (CAS). This technology achieves successful and accurate targeted insertion of large DNA segments in plants.
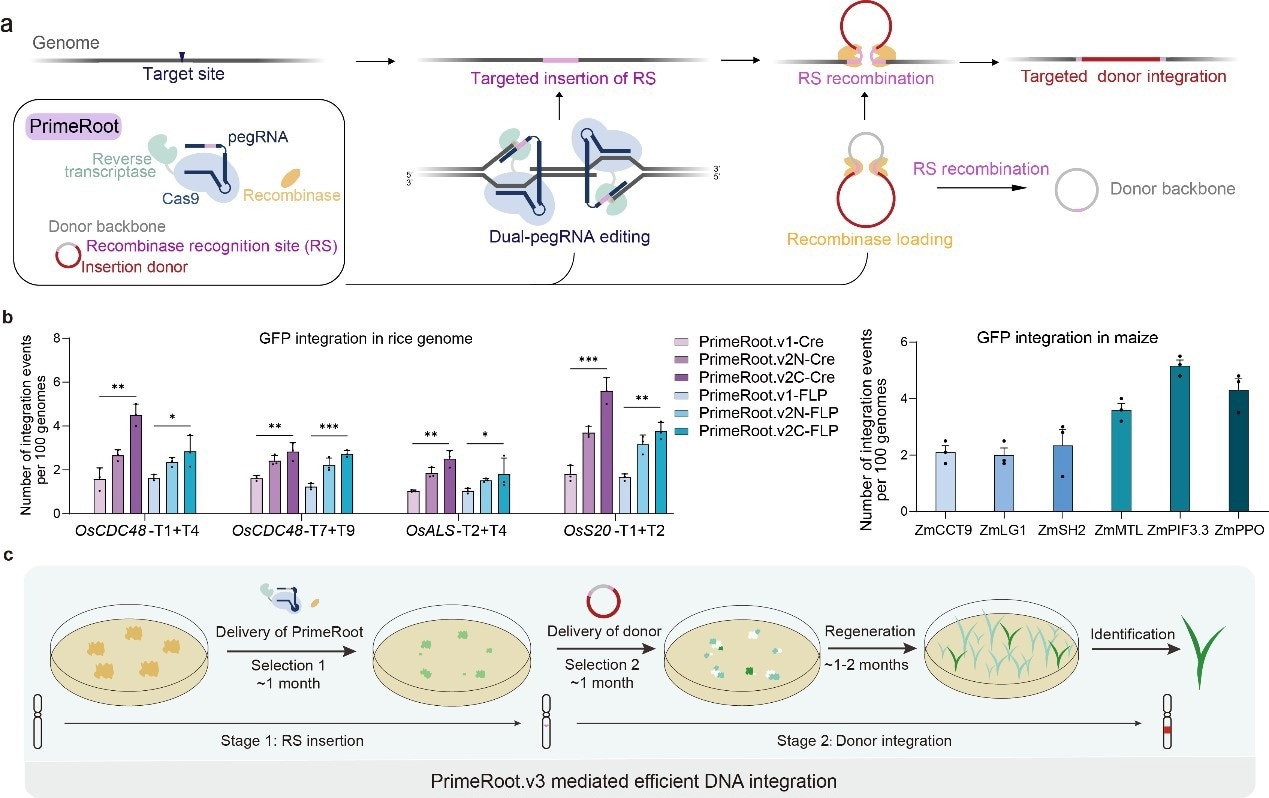
The development and optimization of PrimeRoot editors. a. Schematic overview of the PrimeRoot editor; b. Editing efficiency evaluation of PrimeRoot approaches in rice and maize; c. Schematic overview of efficient PrimeRoot.v3. Image Credit: IGDB
The new technology, known as prime editing-mediated recombination of opportune targets (PrimeRoot), integrates an optimized dual-ePPE editor protein that was earlier published by the team with an extremely successful tyrosine site-specific recombinase, Cre. It can achieve one-step, accurate targeted insertion of large DNA segments in maize and rice with a success rate of a maximum of 6% and has been employed to efficiently insert DNA segments up to 11.1 kb.
Nature Biotechnology published the results on April 24th, 2023.
Genome editing, a disruptive biotechnology, comes with wide effects across the life sciences. In the past 10 years since the first innovative CRISPR-Cas9 studies, the field has progressed from random, intermittent editing to accurate editing technologies. Although prime editing and base editing are successful at making tiny DNA changes, they were unable to edit huge cargoes. As the genome editing field progresses, there is an increasing requirement for targeted, efficient, large DNA insertions into the living cells’ genome.
When compared with the conventional, non-accurate, non-homologous end joining strategy, PrimeRoot has considerably enhanced the efficacy of inserting long DNA segments of 5 kb or more. Mainly, the insertion events are absolutely predictable and accurate.
In this research, the scientists established two particular applications of PrimeRoot editing. PrimeRoot was employed to insert an actin promoter (1.4 kb) upstream of the endogenous OsHPPD gene. The foreign functional elements’ introduction is a significant genetic breeding method to control endogenous gene expression.
Then, PrimeRoot was employed to do targeted gene insertion in plants. Conventional methods founded on Agrobacterium mediation and particle bombardment lead to random and inaccurate insertion events.
To achieve rapid disease resistance breeding, the scientists employed PrimeRoot to precisely insert the rice blast resistance gene pigmR into a predicted genomic safe harbor.
To improve the PrimeRoot efficiency, the scientists demonstrated a sequential transformation system in rice. Then, this system enhanced editing efficacy two to four times in comparison with a single one-shot conversion, thereby gaining maximum efficiencies of 8.3% for inserting an actin promoter (1.4 kb) and efficiencies up to 6.3% for inserting a complete pigmR gene expression frame (4.9 kb).
This technology offers robust technical support for plant synthetic biology research and plant molecular breeding.
Source:
Journal reference:
Sun, C., et al. (2023). Precise integration of large DNA sequences in plant genomes using PrimeRoot editors. Nature Biotechnology, https://doi.org/10.1038/s41587-023-01769-w.