Reviewed by Danielle Ellis, B.Sc.Jun 6 2023
The Hong Kong University of Science and Technology (HKUST) research group, guided by Prof. Tuan Anh Nguyen, Assistant Professor of the Division of Life Science, used several advanced methods, including miRNA sequencing, pri-miRNA structure analysis, and high-throughput pri-miRNA cleavage assays for approximately 260,000 pri-miRNA sequences, to locate and thoroughly illustrate the newly identified noncanonical cleavage mechanism.
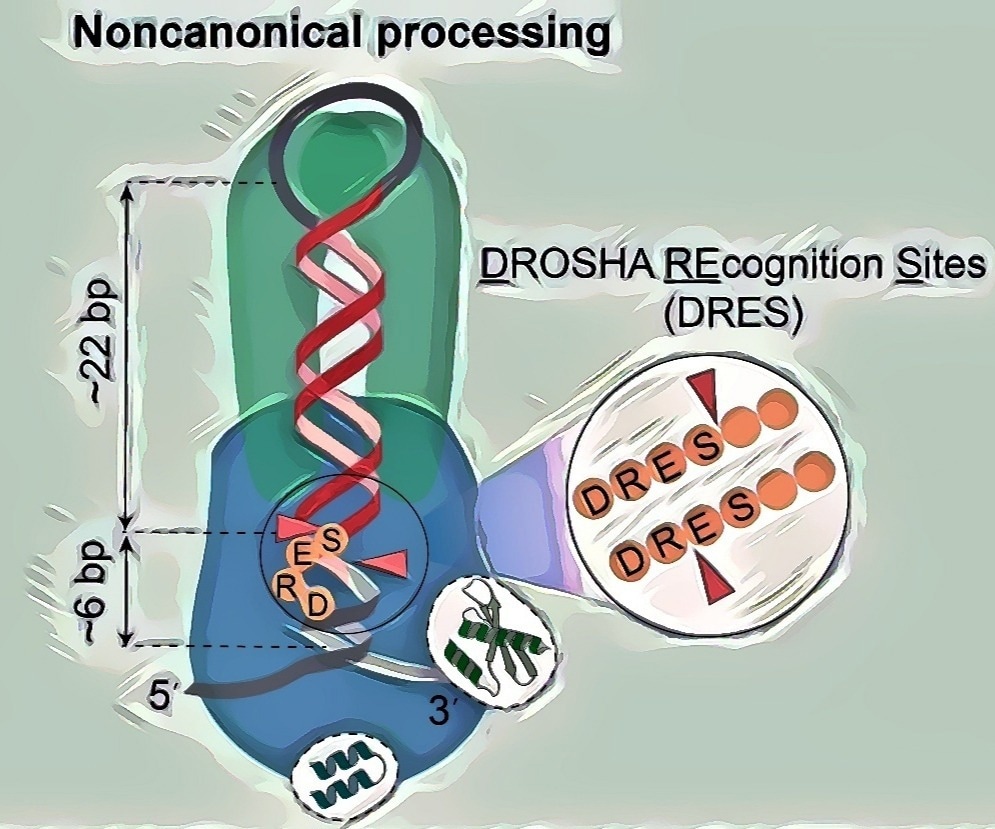 The image depicts the noncanonical processing model of Microprocessors in action. The Microprocessor complex consists of DROSHA, shown in blue, and the DGCR8 dimer, represented in green. Arrowheads indicate the double cleavage action of DROSHA on pri-miRNAs. The noncanonical processing substrate is characterized by a short stem of approximately 28 base pairs and DRES (DROSHA Recognition Sites). This noncanonical mechanism is conserved across a variety of animal species. Image Credit: Prof. Tuan Anh Nguyen.
The image depicts the noncanonical processing model of Microprocessors in action. The Microprocessor complex consists of DROSHA, shown in blue, and the DGCR8 dimer, represented in green. Arrowheads indicate the double cleavage action of DROSHA on pri-miRNAs. The noncanonical processing substrate is characterized by a short stem of approximately 28 base pairs and DRES (DROSHA Recognition Sites). This noncanonical mechanism is conserved across a variety of animal species. Image Credit: Prof. Tuan Anh Nguyen.
The noncanonical process does not rely on certain key protein and RNA elements required by the canonical mechanism.
The research also found previously unknown DROSHA recognition sites (DRES), which are important for noncanonical cleavage but can also act in the canonical cleavage mechanism. The work also emphasizes the evolutionary aspect of this noncanonical cleavage process, indicating that it is conserved across numerous animal species. This discovery implies that the noncanonical mechanism is important in the evolution of miRNA biogenesis and regulation.
MicroRNAs (miRNAs) are small RNA molecules that play an important role in gene regulation. They aid in the regulation of several biological processes, including cell growth, development, and immunity. Scientists have conducted substantial research on miRNAs in recent years to truly comprehend their roles and the mechanisms involved in their production.
Researchers at the Hong Ko

Image Credit: FOTOGRIN/Shutterstock.com
ng University of Science and Technology (HKUST) have made a breakthrough in molecular biology by revealing a noncanonical cleavage mechanism for the Microprocessor (MP, DROSHA-DGCR8 complex) complex, which is responsible for producing miRNAs in humans and other animals by processing primary miRNA transcripts (pri-miRNAs).
This ground-breaking finding provides new light on a long-standing puzzle in molecular biology and has far-reaching ramifications for the knowledge of gene regulation, cellular processes, and the development of miRNA biogenesis pathways in animals.
The Microprocessor complex in animals was identified in 2004, and numerous research groups have examined its molecular mechanisms in generating miRNAs since then. These investigations contributed to the development of a molecular mechanism model of this enzyme known as the canonical pri-miRNA processing mechanism.
This method, however, can only elucidate how the enzyme cleaves numerous pri-miRNAs in animals. Because the structure and sequencing of pri-miRNAs in animals are so variable, this mechanism fails to explain how a major fraction of pri-miRNAs are processed.
The noncanonical pri-miRNA processing mechanism, identified and published in the prestigious journal Molecular Cell, addresses a two-decade-long molecular biology mystery about the cleavage of many pri-miRNAs in animals and supports the previously known canonical mechanism.
In simpler terms, this study discloses a novel method by which human cells generate miRNAs, which could have ramifications for the knowledge of gene regulation, cellular processes, and the development of miRNA biogenesis pathways in animals.
Key Findings from HKUST’s Study
- The identification and complete characterization of a novel method by which cells in animals create miRNAs, thereby resolving a long-standing riddle in molecular biology.
- The discovery of DRES that determine MP cleavage efficiency and precision opens up a new research avenue to investigate whether additional comparable enzymes possess RNA recognition sites.
- Evidence that the noncanonical miRNA synthesis mechanism is conserved across animal species, with worms such as C. elegans and C. briggsae playing an important role in miRNA biogenesis.
- A description of how MP cleaves several short stem pri-miRNAs in animals implies that the MP complex has larger biological activities.
The discovery of the MP complex’s noncanonical cleavage process in miRNA biogenesis has far-reaching ramifications for future molecular biology research. The discovery of this process opens up new research options and broadens the understanding of the regulatory landscape of miRNA biogenesis in animals.
One of the most important implications is the possibility of discovering a wider range of substrates for the MP complex in animals. The canonical mechanism was earlier unable to account for the processing of some pri-miRNAs. Scientists can now examine formerly unexplained or overlooked RNA substrates using the noncanonical method. This could lead to the discovery of new pri-miRNAs and other RNA substrates processed exclusively by the noncanonical mechanism.
Another implication is the possibility of discovering new roles of the MP complex in animals. Because the noncanonical mechanism can process short stem pri-miRNAs, it shows that the MP complex has a larger cellular role. This could pave the way for the discovery of previously undiscovered roles in gene regulation and cellular processes like development, differentiation, and immunity.
Furthermore, by confirming the conservation of the noncanonical mechanism across diverse animal species, particularly in worms such as C. elegans and C. briggsae, the study underscores the mechanism’s evolutionary significance. Future research could explore deeper into the evolutionary elements of both canonical and noncanonical mechanisms, offering light on the development and diversification of animal miRNA biogenesis pathways.
In conclusion, the discovery of the noncanonical mechanism in miRNA biogenesis has broadened the understanding of the molecular mechanisms underpinning miRNA production. This opens the door to future molecular biology discoveries, which will lead to a better understanding of gene regulation, cellular processes, and the development of miRNA biogenesis pathways in animals.
Source:
Journal reference:
Nguyen, T. L. (2023). Noncanonical processing by animal Microprocessor. Molecular Cell. doi.org/10.1016/j.molcel.2023.05.004.