Unsafe foods have debilitating human, economic and social consequences.
Salmonella is 1 of 4 key global causes of diarrheal diseases (WHO), causing about 1.35 million infections and 420 deaths in the United States only every year (US CDC).
Contamination of food with harmful bacteria, such as Salmonella, makes food unsafe. Salmonella can affect the health of anyone who eats or handles contaminated foods, and if consumed can cause illness, commonly known as food poisoning.
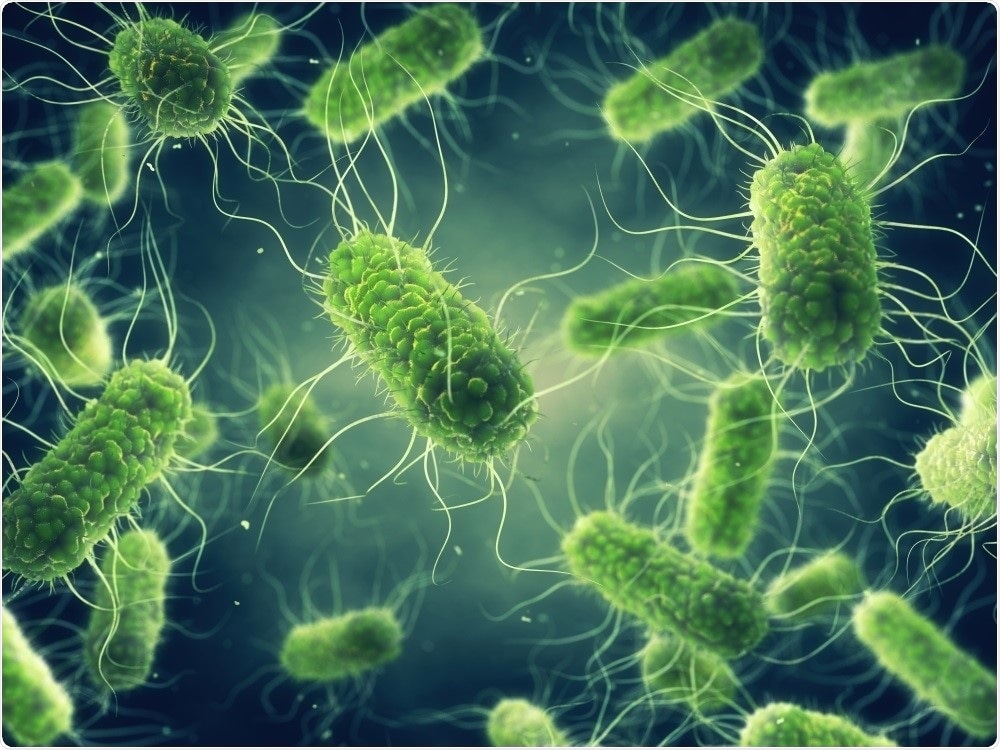
Image Credit: nobeastsofierce/Shutterstock.com
At Mars, we believe everyone has the right to safe food and as a global food manufacturer, we believe we have a clear responsibility to lead in food safety. However, no single entity can do this alone. Collaboration is critical.
That is why, in September 2015, we opened the Mars Global Food Safety Center (GFSC). At the center, we conduct original research and collaborate to exchange knowledge and insights in food safety to help ensure safe food for all.
One of the areas we focus on with that is microbial risk management and Salmonella prevention and control is a key area of research.
Not only is microbial risk management an area of great importance for the food industry, but it is also critical for ensuring safe food. There is a significant opportunity to increase understanding of the behavior of bacteria or pathogens in food environments and enhance food safety management.
How does salmonella occur within food?
Salmonella can naturally occur in the gut of many animals, including wild animals, farm animals, and pets. As it is associated with animals, it is not uncommon for it to disperse.
The best approach is to always treat food as if it is contaminated and there are clear ways to do that in food handling. When preparing meat, for example, it is important to ensure that everything is cooked thoroughly.
Heating to a minimum temperature of 160 Fahrenheit or 70 degrees Celsius is usually sufficient to eliminate Salmonella.
At the same time, it is important to guard against cross-contamination of different (raw, cooked) meats and ensure that raw and cooked meat are not chopped on the same chopping board.
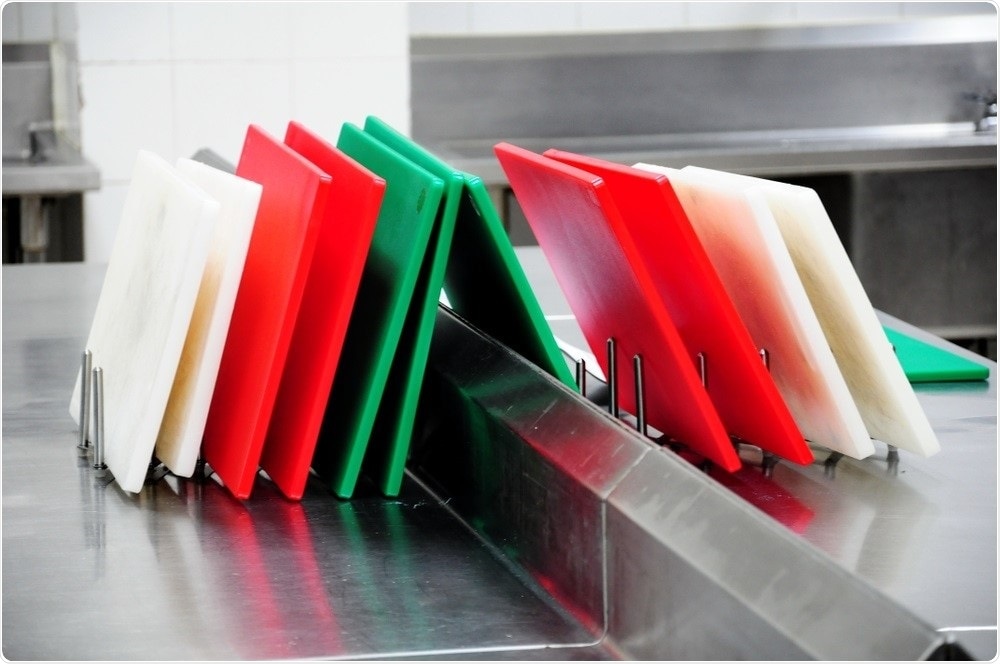

Image Credit: Tarn Tiny/Shutterstock.com
While safe food handling, preparation, and storage can help reduce the risk of Salmonella and cooking thoroughly further reduces the risk Salmonella, food can be contaminated after processing due to cross-contamination through contact with other food materials.
This is the reason that food manufacturers have rigorous and robust quality and food safety risk management processes in place. Food manufacturers rely on proper segregation and the application of the right processes to minimize the risk of Salmonella contamination.
How do we currently characterize salmonella?
Characterization is important in source tracking. When embarking on the characterization of Salmonella it is important to start with the genus (Salmonella), and from there work to understand the specific family that the strain belongs to.
Characterizing the strain helps with understanding the root cause of a specific Salmonella outbreak enabling investigation of where it came from, how it got there and then taking steps to eradicate it.
Conventional ‘serotyping’ (identification of the type of Salmonella) has been used as a Salmonella subtyping method for more than 80 years and has been a certified approach for public health monitoring of Salmonella infections for over 50 years.
While it is robust and reliable, the conventional route is also time-consuming (usually taking from three to more than 12 days) and labor-intensive.
Molecular-based subtyping methods for Salmonella identification have been developed to replace the conventional serotyping method. These methods generally use direct targets (genetic determinants of serotypes) or indirect targets (surrogate genomic markers associated with particular serotypes) for serotype determination.
Some of these molecular-based subtyping methods are less effective at differentiating genetically closely related serotypes and are sometimes limited by relatively small database sizes when predicting Salmonella serotypes. These limitations can be overcome by the use of Whole Genome Sequencing (WGS)-based serotyping methods.
One of the goals of our study was to understand if we could improve the efficiency of strain identification in order to increase the ability to act faster. In the study, entitled: ‘Evaluation of Real-Time Nanopore Sequencing for Salmonella Serotype Prediction’, we worked together with the University of Georgia and Cornell University to explore the possibilities of new technology in helping to open new solutions.
What we found was that using this method, completion of WGS and corresponding genetic serotyping is possible within eight hours - much faster than conventional methods.
Further work is needed but this development could improve the speed and accuracy of pathogen identification which has implications for the food industry and the global food supply chain as a whole.

Image Credit: KAMONRAT/Shutterstock.com
Why is it important to be able to rapidly and accurately track pathogen contaminations within food?
Having the ability to identify strains quickly and act fast is critical in efforts to ensure safe food.
As the food supply chain becomes ever more global and interconnected, the opportunity for food to become contaminated increases.
In the fast-moving world of food manufacturing, where rapid identification and response to Salmonella contamination incidents is critical, developing a more efficient pathogen identification method is essential.
The quicker the source of contamination can be identified, the quicker the contamination can be eradicated.
More rapid and accurate tracking of the pathogen can help to find the root cause of Salmonella contamination along the food supply chain faster and more precisely.
What is Whole-Genome Sequencing (WGS) and how is this applied to characterize salmonella?
The long turnaround times, high costs and complex sample preparations of previous methods mentioned have led global food safety regulators, food authorities and public health agencies to change to WGS methods for pathogen subtyping.
WGS is a method used to capture and analyze all the information of the entire genome of an organism in one process. It provides a very precise DNA fingerprint that can help link foodborne pathogen cases to one another, allowing an outbreak to be detected and solved sooner.
Through the Mars GFSC, we are also actively exploring and evaluating new, innovative methods and capabilities such as whole-genome sequencing with nanopore long-read sequencing technologies, aiming to continuously improve the food industry approach to food safety.
The WGS-based serotyping method of our study directly uses serotype determinant DNA sequences for the serotype prediction of Salmonella. The nanopore sequencing technology we used for this allows the sequencing run to be stopped when sufficient output has been generated for serotype prediction.
Our method, therefore, needs as little as two hours upwards of WGS, which can enable completion of the whole prediction process within 8 hours.
In your research, you mention that there are different subtypes of salmonella. Can you describe the differences between these subtypes?
Serotyping classifies the genus Salmonella into serovars (also known as “serotypes”) based on surface antigens – characteristics in a way.
In all, more than 2,500 serovars of Salmonella enterica - the Salmonella species responsible for virtually all salmonellosis cases - have been identified using conventional serotyping, but less than 100 serovars account for the vast majority of human infections (CDC 2015).
These differences provide insights that enable identification and therefore the opportunity to isolate and eradicate them.
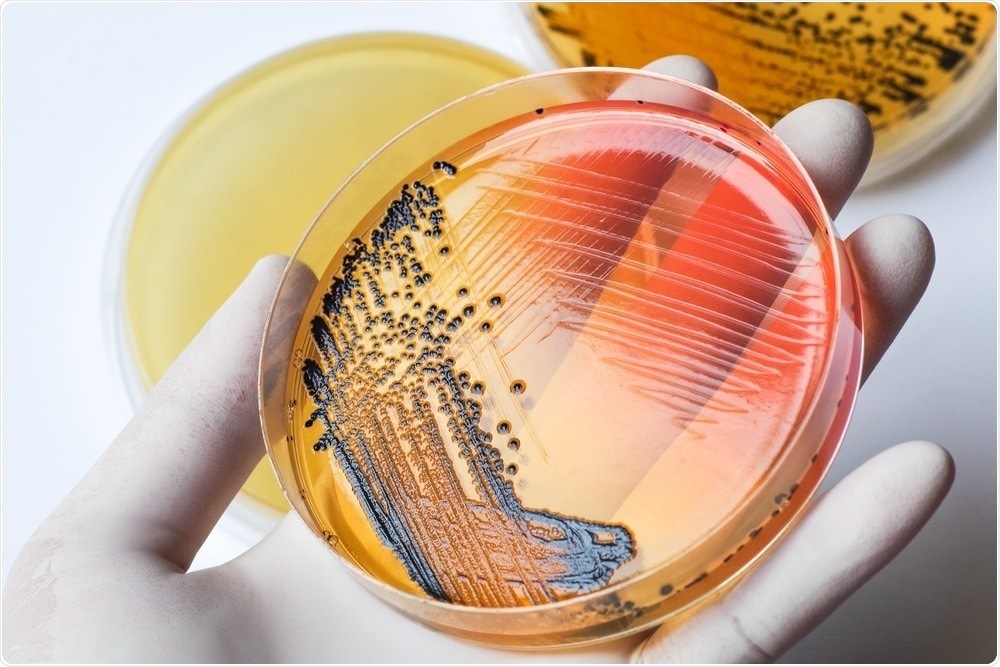
Image Credit: Chansom Patnip/Shutterstock.com
What are the drawbacks of using WGS within the food industry?
The accurate identification of Salmonella serotypes is critical and the application of DNA-based methods for identification and characterization of pathogens such as Salmonella has become common practice.
Looking at different options for this, after comparing resolution, turnaround time, ability to predict serovars, cost and feasibility of different DNA-based methods, WGS in my view, is the most suitable method to identify and characterize Salmonella in response to incident investigation for the production facilities in the food industry.
As next-generation sequencing technologies are continually developed to allow for more rapid, user-friendly and cost-effective outcomes, WGS is increasingly being adopted as a pathogen subtyping tool and beyond as next-generation sequencing platforms can be widely applied for multiple purposes.
The genomic sequencing information can be used to monitor ingredient supplies, antimicrobial resistance and persistency of a pathogen (including Salmonella) in a processing environment, as well as facilitate the improvement of sanitary management, microbial hazard control, and microbiological risk assessment, to name a few.
These applications of WGS will help the food industry shift from being reactive to incidents caused by microbial contamination to proactive. With the full application of WGS, it will be possible to achieve a well monitored and microbial risk-controlled food supply chain that ensures safe food for all.
Nevertheless, there is a strong need at the global level for the standardization of WGS operation and data analysis, in particular, source tracking analysis.
A common agreement of understanding and applying WGS among the food industry and authorities from food safety and public health is expected to guide the implementation of WGS in food safety management.
What other methods could the food industry use to overcome these issues?
In the long run, metagenomics which also uses genomics technology may become an even more powerful tool than WGS for the food industry.
One collaborative project which involves metagenomics is the Consortium for Sequencing the Food Supply Chain (CSFSC) - established in 2015 by IBM and Mars, Incorporated to help drive global innovation in genomics, food, and agriculture, enabling critical breakthroughs in food safety.
The consortium aims to take food safety to a new level through the application of genomics and big data that will generate new insights and understanding of the total supply chain, enabling the controls and standards to be evolved and applied in new ways.
Work has been completed to validate the initial hypothesis - that the microbiome could be useful for food safety - as unique and diverse microbiological indicators have been found in a limited number of raw materials.
A baseline has been observed for samples of similar matrix composition and the microbiome can signal outlier samples in the supply chain, i.e. potential contaminants.
This is an important finding in the context of food safety management and will require extensive data at a much bigger scale to further validate the science and complete the picture for the food supply chains managed around the world.
What are the next steps in your research into salmonella?
The study we carried out is a proof-of-concept study to evaluate Oxford Nanopore Technologies’ (ONT) long-read sequencing platform for Salmonella serotype prediction in the arena of food safety, together with the University of Georgia and Cornell University.
As always, there is more we wish to do but this study provides a starting point for future validation of ONT sequencers for Salmonella subtyping on a more extensive strain set and their potential application in the food industry for Salmonella contamination source tracking.
At Mars, we believe that everyone has the right to access safe food and that as a global manufacturer that we have a responsibility to take the lead in sharing knowledge and insights.
Research such as the Evaluation of Real-Time Nanopore Sequencing for Salmonella Serotype Prediction”, forms an important of our commitment to exploring and applying ways to help ensure safe food for all.
As part of our Microbial Risk Management program at the Mars GFSC, it forms an important part of our work dedicated to applying cutting edge technology and solutions to raise the bar in microbial food safety and help create a better world tomorrow.
Where can readers find more information?
About Dr. Silin Tang
Silin Tang is a Senior Research Scientist at the Mars Global Food Safety Center, where Silin leads research projects on the development, evaluation, and validation of pathogen detection and identification and subtyping methods for the food industry, including the application of whole-genome sequencing technologies in source tracking of foodborne pathogens.
Silin graduated from Cornell University with a Ph.D. degree in Food Science and Technology and has more than 10 years of research experience working in microbiological food safety.
Her research focus at Cornell included the development of control strategy against foodborne pathogens across the food supply chain, transcriptome analysis of foodborne pathogens under stress conditions, and the application of sequencing technologies in bacteria subtyping.
Silin has also worked at the Clinic for Poultry, University of Veterinary Medicine, Hannover, Germany, and at BioMeireux in St. Louis, USA.