In the field of structural biology, certain molecules are so uncommon that they can only be captured with a special set of tools.
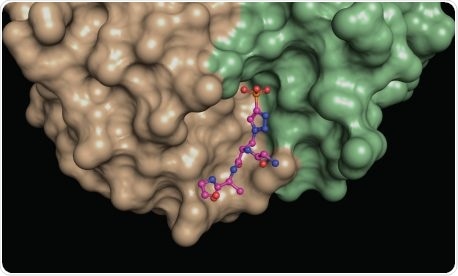
A structural snapshot of a phosphohistidine analog (ball and stick model) nestled at the interface between different areas (green, brown) of a phosphohistidine antibody. Such structures provide insights into the molecular properties of the antibodies, which makes them useful for revealing elusive phosphohistidine-containing proteins in cells. Image Credit: Salk Institute.
That is exactly how a multi-institutional research team, headed by scientists from Salk Institute, defined how a compound known as phosphohistidine can be detected by antibodies. Phosphohistidine is an extremely unstable molecule that has been known to play a crucial role in certain forms of cancer, like neuroblastoma and breast and liver cancer.
These latest insights will prepare the research team for more sophisticated analyses on phosphohistidine and its promising role in cancer, and will also allow them to exploit the shape and atomic composition of the binding sites of the antibodies to engineer even more efficient antibodies in the days to come.
The new study was published in the Proceedings of the National Academy of Sciences on February 5th, 2021.
We are excited that these new antibody structures reveal novel principles of antigen binding. Now we can redesign these antibodies and engineer their properties to make them more efficient. This work may also provide other scientists with phosphohistidine antibodies that better suit their research purposes.”
Tony Hunter, Renato Dulbecco Chair and American Cancer Society Professor, Salk University
Hunter is also the senior author of the study.
Amino acids are combined together in accurate sequences to create proteins, and many of them can go through chemical changes that can more or less alter the protein activity. One such chemical change is a process known as phosphorylation—when a compound, known as phosphate, is added to an amino acid, altering its charge and shape.
Hunter has previously demonstrated that phosphorylation on the amino acid tyrosine can fuel the progression of cancer, a finding that resulted in various anticancer drugs. More lately, Hunter focused on phosphorylation of the amino acid, called histidine (which produces phosphohistidine), assuming that the process may also play a crucial role in human diseases.
Hunter designed a set of antibodies that can attach to phosphohistidine in proteins, and utilized chemically stabilized phosphohistidine analogs to create a suite of monoclonal antibodies that could detect these forms.
The following step was to precisely interpret how the antibodies can attach to phosphohistidine. This made Hunter team up with Ian Wilson, the Hansen Professor of Structural Biology at the Scripps Research Institute and a famous expert in using protein crystallography to characterize the structures of antibodies, to investigate the structures of the phosphohistidine antibodies.
My long-term colleague Tony and I have been collaborating on this project for the past seven years. We have obtained new insights into how antibodies can evolve to recognize phosphates linked to proteins, which is very satisfying.”
Ian Wilson, Hansen Professor of Structural Biology, Scripps Research Institute
To determine how phosphohistidine is detected, the duo had to image the antibodies of phosphohistidine during the act of binding the phosphohistidine and thus formed crystals between every antibody attached to a phosphohistidine peptide.
To understand the molecular interactions between the antibodies and phosphohistidine, we needed to look at them in great detail.”
Rajasree Kalagiri, Study First Author and Postdoctoral Researcher, Salk University
Kalagiri, who is also an expert in X-ray crystallography, continued, “Once we got the antibodies to form crystals, we bombarded those crystals with X-rays to obtain a diffraction pattern. We then applied methods that transform the diffraction pattern into a three-dimensional electron density map, which was then used to discern the atomic structure of the antibodies.”
Both types of antibody crystal structures resolved by the researchers precisely demonstrated how different types of amino acids are organized around the phosphohistidine to bind it firmly. The five structures of antibodies more than doubled the number of phospho-specific antibody structures reported before, and this gives new insights into how antibodies detect both the phosphate and the linked histidine.
The team also demonstrated at a structural level how both types of antibodies detected diverse forms of phosphohistidine, and this understanding will enable the researchers to design enhanced antibodies in the days to come.
Source:
Journal reference:
Kalagiri, R., et al. (2021) Structural basis for differential recognition of phosphohistidine-containing peptides by 1-pHis and 3-pHis monoclonal antibodies. Proceedings of National Academy of Sciences. doi.org/10.1073/pnas.2010644118.