Alzheimer’s disease (AD) is a neurological condition that leads to dementia worsening with age as patients show memory, cognitive, and psychological deficits. Existing therapies concentrate on easing these symptoms to some extent.
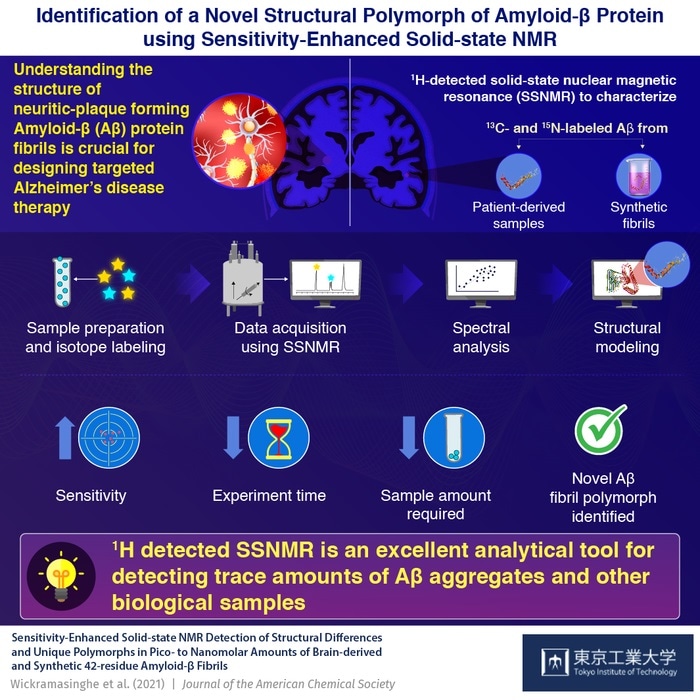
Sensitivity-enhanced solid-state NMR spectroscopy can be used in the structural characterization of amyloid β—the pathogenic protein involved in Alzheimer’s disease—as shown by scientists from Tokyo Tech. Their findings bring to light a novel polymorph of the protein and associated structural elements that can be targeted for disruption towards the development of new AD treatment strategies. Image Credit: Tokyo Tech.
AD does not have a definite cure or prevention, implying the need for continued efforts in gaining better knowledge on the biology of the disease.
Pathogenic amyloid β (Aβ) protein fibrils are accumulated in the form of plaques in the brain in AD. Hence, deciphering its structural organization is vital for developing targeted treatments against the disease.
Even though numerous forms, or polymorphs, of Aβ40 species of the protein, are reported, there is a vague knowledge about the more pathogenic species, Aβ42. Also, characterizing trace quantities of Aβ from a small sample amount employing standard analytical techniques yet remains a challenge.
A group of scientists from Tokyo Institute of Technology, RIKEN, the University of Illinois at Chicago, and the University of Chicago, headed by Prof Yoshitaka Ishii, analyzed the applicability of solid-state nuclear magnetic resonance (SSNMR) spectroscopy in decoding the atomic-level structural differences of Aβ and associated pathogenic fibrils.
This robust analytical technique measures the differential behavior and properties of nuclei under the influence of magnetic and electric fields, emphasizing their atomic structure. The research outcomes have been published in the Journal of the American Chemical Society.
Limited information is available on structural variations of Aβ42 fibrils prepared at physiologically relevant conditions, despite their pathological importance. In our study, we demonstrate the use of 1H (hydrogen isotope)-detected SSNMR in the characterization of patient-derived, as well as synthetic Aβ fibrils in limited amounts as low as pico- to nano-moles.”
Yoshitaka Ishii, Professor, Tokyo Institute of Technology
13C (carbon isotope)-detected SSNMR, conventionally utilized for structural characterization, involves a large sample amount and raises difficulties in the preparation of homogenous samples. The researchers employed sensitivity enhanced 1H detected SSNMR for their analysis, provided the higher sensitivity and feasibility of analyzing trace amounts in biological samples.
Synthetic fibrils and brain-derived Aβ from a patient with Alzheimer’s disease were labeled with isotopes 13C and 15N at specific amino acid residues for atomic resolution, enhanced sensitivity, and site-specific structural analysis.
The researchers efficiently characterized a novel polymorph of Aβ42 using the above-mentioned process, employing only about 42 nmol of Aβ—25 to 100 times less than earlier used concentrations. Moreover, sensitivity enhancement substantially decreased the time required to get the spectra of the samples.
Spectral positions obtained by this approach disclosed that the structure of the protein backbone and the arrangement of the side chains are different from earlier reported structures.
The research explains the molecular contacts involved in stabilizing pathogenic Aβ42 fibrils, thereby leading the path for novel therapeutic approaches with the ability to target these toxic aggregates that drive AD progression.
Our study demonstrates the propensity of Aβ42 to form multiple forms of fibrils, in the brain as well as in synthetic preparations. We believe that our study can open new avenues for analyzing trace amounts of biological samples such as amyloid fibrils and oligomers, for which 13C-detected SSNMR might be ineffective.”
Yoshitaka Ishii, Professor, Tokyo Institute of Technology
The research findings help set a foot forward in the understanding of this complex disease.
Source:
Journal reference:
Wickramasinghe, A., et al. (2021) Sensitivity-Enhanced Solid-State NMR Detection of Structural Differences and Unique Polymorphs in Pico- to Nanomolar Amounts of Brain-Derived and Synthetic 42-Residue Amyloid-β Fibrils. Journal of the American Chemical Society. doi.org/10.1021/jacs.1c03346.