Scientists from Japan recently designed the first bottom-up designed peptides, containing chains of amino acids, that can develop into artificial nanopores to pinpoint and allow single molecule-sorting of genetic material in a lipid membrane.
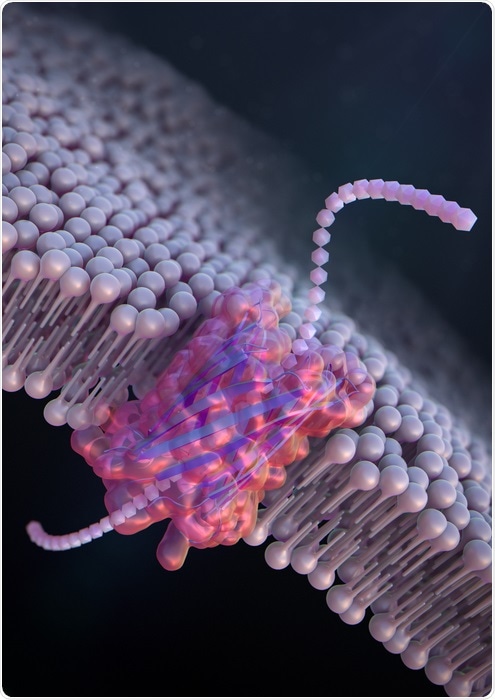
De novo design of a nanopore for single-molecule detection that incorporates a β-hairpin peptide. Image Credit: Ryuji Kawano, Tokyo University of Agriculture and Technology.
Biological nanopores are channels created by pore-forming proteins, that can identify certain molecules, however, such natural channels are hard to detect, restricting proposed applications in inexpensive, small molecule identification, speedy DNA sequencing, and much more.
Nanopore sensing is a powerful tool for label-free, single-molecule detection. This is the first time that DNA and polypeptides were sensed using a de novo-designed nanopore.”
Ryuji Kawano, Corresponding Author, Professor, Tokyo University of Agriculture and Technology
The observations were published on November 22nd, 2021, in the Nature Nanotechnology journal.
According to Kawano, the de novo-designed nanopores are created “from scratch” and have the capability to imitate natural proteins and their capability to identify specific proteins. Kawano added that they could be engineered to function as artificial molecular machines capable of identifying a much broader range of molecules—which might help explain the link between structure and function in target proteins.
The folded structure of proteins is determined by their linear polypeptide sequence and gives rise to specific protein functionality.”
Ryuji Kawano, Corresponding Author, Professor, Tokyo University of Agriculture and Technology
Kawano remarked noting that all proteins have an exceptional structure and size.
The unique primary structure is the result of structural evolution such as the mutation and selection of amino acid residues over time. To reveal the relationship between this primary information and protein structure is one of the ultimate goals of science.”
Ryuji Kawano, Corresponding Author, Professor, Tokyo University of Agriculture and Technology
Kawano and the group designed a peptide dubbed SV28 to create huge synthetic nanopores that enable better detection and identification of molecules for practical applications. By bending the two arms of amino acids at a sharp angle, and with specific charges at the terminus.
By applying a voltage, the orientation of the hairpin-shaped peptide can be controlled precisely. The peptide can assemble to form nanopore structures in the size range of 1.7 to 6.3 nm, appropriate for identifying molecules of DNA.
The scientists also modified SV28 by introducing a mutation that prompts the peptide structure to bend and twist in certain ways. The resultant peptide developed evenly dispersed pores of 1.7 nm each, able to detect a single polypeptide chain—or one-half of a protein.
This accomplishment can be applied to promote the understanding of the relationship between protein structure and function.
As a further step, the group intends to design different peptides and proteins to build various types of nanopores to help in peptide sequencing, function as molecular robots, and more.
Source:
Journal reference:
Shimizu, K., et al. (2021) De novo design of a nanopore for single-molecule detection that incorporates a β-hairpin peptide. Nature Nanotechnology. doi.org/10.1038/s41565-021-01008-w.