Researchers from the Scripps Institution of Oceanography at UC San Diego, the J. Craig Venter Institute (JCVI), and the National Oceanic and Atmospheric Administration (NOAA) evaluated the diversity of aquatic life off the California coast using genetic research tools similar to those used in genealogy.
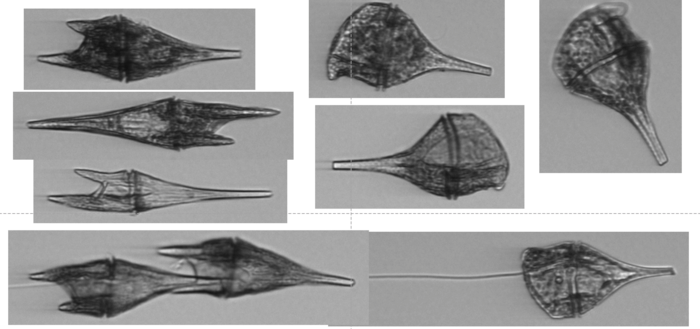 Dinoflagellates collected during NSF Long-term Ecological Research cruise, 2021. Image Credit: Andrew Allen
Dinoflagellates collected during NSF Long-term Ecological Research cruise, 2021. Image Credit: Andrew Allen
Researchers will be able to utilize the technology to identify abnormalities at the base of the ocean food web that affect the quantity of commercially valuable fishes or cause toxic algal blooms. Researchers can utilize so-called environmental DNA (eDNA) to assess how well the oceans can buffer the globe from the effects of climate change based on data acquired through a method called “metabarcoding.”
The results were published in the journal Nature Communications on May 4, 2022. The National Science Foundation (via the California Current Ecosystem Long-Term Ecological Research project), NOAA, and the Gordon and Betty Moore Foundation supported the study.
It’s the ecological sampling method of the future. This study represents the first deployment of this approach within a long-term ecological sampling context. It reveals what you can see when all this hidden diversity is finally shown.”
Chase James, Study First Author and Graduate Student, Scripps Institution of Oceanography
James was also a researcher at JCVI.
The new method of evaluating ocean microbiomes—collections of microscopic plants, animals, and other species living in specific habitats—substantially increases scientists’ capability to do ocean diagnostics.
In this research, scientists were able to discover the most critical element determining how many species are in the ocean in surface waters off the California coast and where they are distributed using genetic information. They discovered that nutrition supply influences the microbial life profile in the California Current more than temperature. This result could not have been reached through traditional methods.
The technique, according to James, is similar to scanning the barcodes of all the products at a grocery shop to get an inventory of them. The NOAA CalCOFI Ocean Genomics Project (NCOG), led by James’ adviser Andrew Allen, began in 2014 with water samples collected during cruises of the renowned CalCOFI surveys, a quarterly program that Scripps has co-managed since 1949.
Filtered samples were obtained in two-liter bottles, and the filters were frozen and sent to the lab. The researchers then identified all of the microbes in the samples by profiling all of the DNA they found in them in the same way that commercial DNA testing businesses identify people’s genetic profiles. They also calculated the number of specimens of each of the identified species in the sample.
Traditional approaches such as light microscopy, which captures sentinel species often found in saltwater, or bulk indicator readings such as how much chlorophyll is in the water, have been improved by the process. Those technologies, in comparison to metabarcoding, only provide broad strokes-level information about where life lives. Metabarcoding enables for more precise species identification and data collecting with the same amount of work.
CalCOFI was established shortly after WWII to assist officials and the fishing sector in determining what caused the dramatic decline of sardine numbers off the West Coast. Quarterly cruises are held at a variety of places off the coast as part of the program. Researchers repeat a series of physical and biogeochemical tests to determine ecological conditions. Scientists have compiled an unrivalled history of the maritime environment because of the surveys.
It’s interesting that 70 years ago, CalCOFI couldn’t have even imagined that you could sample two liters of seawater and get comprehensive data on the marine microbial community, but a major future goal of this study is to achieve the initial goals that CalCOFI set out to accomplish, which is to understand the processes that drive the success and failure of our regional fisheries. This cutting-edge research may be used to answer 70-year-old questions.”
Chase James, Study First Author and Graduate Student, Scripps Institution of Oceanography
Source:
Journal reference:
James, C. C., et al. (2022) Influence of nutrient supply on plankton microbiome biodiversity and distribution in a coastal upwelling region. Nature Communications. doi.org/10.1038/s41467-022-30139-4.