Customized protein design is now possible, thanks to artificial intelligence (AI), which can be used to address both medicinal and environmental issues. Now, a computer-based natural language processing model has been effectively used for protein research by a team at the University of Bayreuth under the direction of Prof. Dr Birte Höcker.
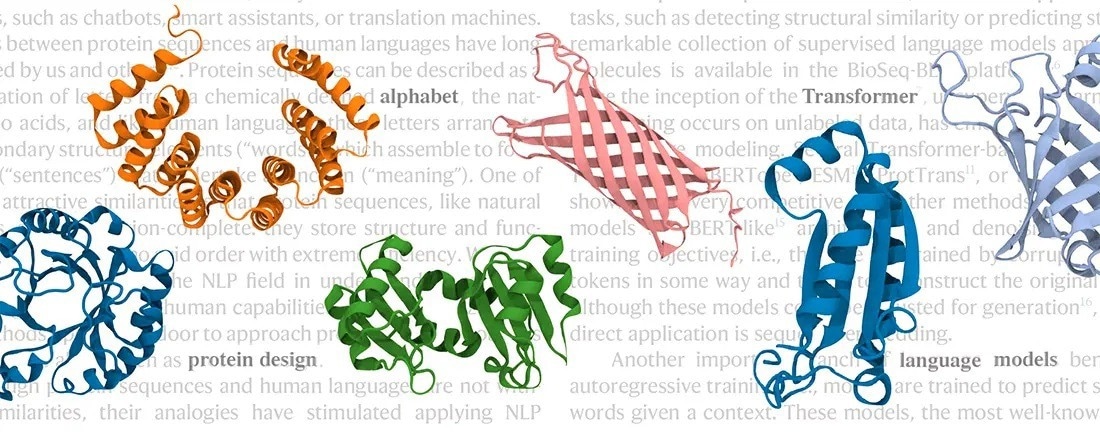 Principles and processes that govern computational natural language processing are now increasingly used in protein research. Image Credit: University of Bayreuth /Protein design group.
Principles and processes that govern computational natural language processing are now increasingly used in protein research. Image Credit: University of Bayreuth /Protein design group.
The ProtGPT2 model creates new proteins independently that are capable of stable folding and have the potential to take over specific roles in more complex molecular environments. In the journal “Nature Communications,” the model and its prospects are described in detail.
Proteins and natural languages actually have similar structural features. Amino acids arrange themselves into a variety of combinations to build structures that serve specific roles in the living organism, similar to how words construct sentences in many combinations to express specific facts.
As a result, a variety of strategies have been created recently to apply concepts and procedures that govern the computer-assisted processing of natural language in the field of protein research.
Natural language processing has made extraordinary progress thanks to new AI technologies. Today, models of language processing enable machines not only to understand meaningful sentences but also to generate them themselves. Such a model was the starting point of our research. With detailed information concerning about 50 million sequences of natural proteins, my colleague Noelia Ferruz trained the model and enabled it to generate protein sequences on its own.”
Dr Birte Höcker, Professor and Head, Protein Design Group, University of Bayreuth
Dr Birte Höcker adds, “It now understands the language of proteins and can use it creatively. We have found that these creative designs follow the basic principles of natural proteins.”
“ProtGPT2” is the name of the language processing model that was applied to protein evolution. It can now be used to create proteins that fold into stable structures and remain functional in this condition indefinitely. Through extensive research, the Bayreuth biochemists have also discovered that the model is capable of producing proteins that do not exist in nature and may not have ever existed in the course of evolution.
These discoveries open the door to constructing proteins in unique and uncharted ways and provide insight into the infinite universe of potential proteins. Another benefit is that the majority of proteins created from scratch so far have idealized structures.
Such structures typically go through a complex functionalization process before they may potentially be used, such as introducing extensions and cavities, to interact with their surroundings and take on precisely defined functions in larger system settings. On the other hand, ProtGPT2 produces proteins that are already functional in their respective contexts and have such distinct architectures naturally.
Our new model is another impressive demonstration of the systemic affinity of protein design and natural language processing. Artificial intelligence opens up highly interesting and promising possibilities to use methods of language processing for the production of customized proteins. At the University of Bayreuth, we hope to contribute in this way to developing innovative solutions for biomedical, pharmaceutical, and ecological problems.”
Dr Birte Höcker, Professor and Head, Protein Design Group, University of Bayreuth
Source:
Journal reference:
Ferruz, N., et al. (2022) ProtGPT2 is a deep unsupervised language model for protein design. Nature Communications. doi.org/10.1038/s41467-022-32007-7.