The recently released cancer protein profile database, which was created using AI and machine learning, has helped with cancer prediction medicine. The new open-access Disease Blood Atlas, unveiled by KTH Royal Institute of Technology Professor Mathias Uhlén, gives the first-ever map of the proteome signature in cancer patients’ blood.
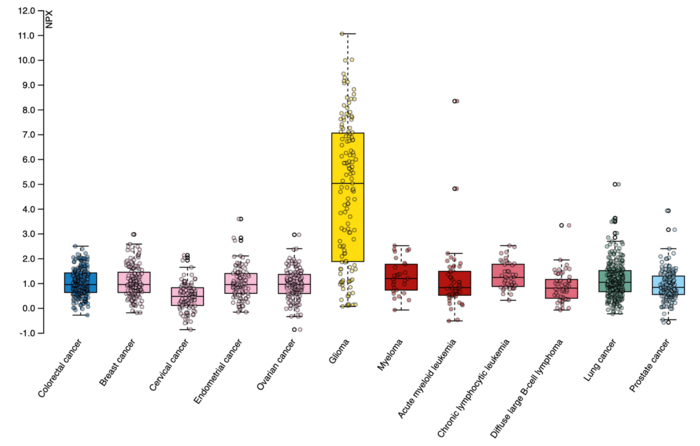 The yellow bar in this readout of protein GFAP indicates elevated expression in the blood of patients with brain tumors. Image Credit: Human Disease Blood Atlas.
The yellow bar in this readout of protein GFAP indicates elevated expression in the blood of patients with brain tumors. Image Credit: Human Disease Blood Atlas.
The Disease Blood Atlas shows 1,463 proteins linked to 12 different types of cancer, as well as proteins that can be used to accurately identify cancer types from a drop of blood.
The Human Protein Atlas consortium, coordinated by Uhlén and based within SciLifeLab, a collaborative research facility that includes KTH Royal Institute of Technology, Uppsala University, Karolinska Institutet, and Stockholm University, created the new database.
The Disease Blood Atlas, according to Uhlén, was constructed using measurements of minute amounts of blood plasma obtained from 1,400 cancer patients at the time of diagnosis and prior to treatment. The blood samples were subjected to a mix of statistical gene expression analysis and machine-learning-based illness prediction.
This is a novel pan-cancer strategy for exploring the proteome signature in blood from cancer patients.”
Mathias Uhlén, Professor, KTH Royal Institute of Technology
The 22nd iteration of the open-access Human Protein Atlas, a resource for profiling human proteins, comprises 12 parts each studying human proteins from different angles, including the new Disease Blood Atlas and Protein 3-D Structure sections.
The release is accompanied by five million pages of revisions to the Human Protein Atlas’ tissue and cell line databases.
The Protein 3-D Structure section displays the 3-D structures of all human proteins based on an AI-based prediction model (AlfaFold). Furthermore, a significant enhancement to the Tissue Atlas section includes precise multiplex spatial profiling of proteins specific to the human testis and kidney. More information on single cell examination of tissues and organs, as well as data from a large collection of human cell lines, is also supplied.
We believe that the new sections of the open access Human Protein Atlas with large amounts of novel data covering all human proteins provides new dimensions of valuable information for researchers interested in human biology and disease.”
Mathias Uhlén, Professor, KTH Royal Institute of Technology