Reviewed by Danielle Ellis, B.Sc.Jul 19 2023
High-brilliance X-Rays from BESSY II can be utilized to create microscopic images with a spatial resolution of as little as a few tens of nanometers. Without the complicated sample preparation required for electron microscopy, whole cell volumes can be analyzed.
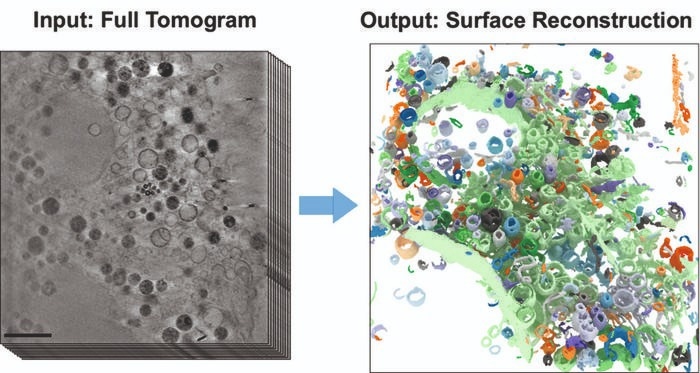 The images show part of a frozen mammalian cell. On the left is a section from the 3D X-Ray tomogram (scale: 2 μm). The right figure shows the reconstructed cell volume after applying the new AI-supported algorithm. Image Credit: HZB
The images show part of a frozen mammalian cell. On the left is a section from the 3D X-Ray tomogram (scale: 2 μm). The right figure shows the reconstructed cell volume after applying the new AI-supported algorithm. Image Credit: HZB
The microscopic cell organelles, with their intricate structures and boundary membranes, are visible under an X-Ray microscope in great clarity and detail, even in three dimensions.
This makes cryo X-Ray tomography suitable for examining changes in cell structures produced by extrinsic events, for example. However, until now, evaluating 3D tomograms required mostly manual and labor-intensive data analysis.
Teams led by computer scientist Prof. Dr Frank Noé and cell biologist Prof. Dr Helge Ewers (both from Freie Universität Berlin) have now partnered with the HZB’s X-Ray microscopy department to address this challenge.
Image Credit: Meletios Verras/Shutterstock.com
The computer science group created a revolutionary self-learning algorithm. This AI-based analysis technique speeds the quantitative analysis of 3D X-Ray data sets by automating the discovery of subcellular structures. The 3D images of the interior of biological samples were obtained at BESSY II’s U41 beamline.
In this study, we have now shown how well the AI-based analysis of cell volumes works, using mammalian cells from cell cultures that have so-called filopodia.”
Dr Stephan Werner, Helmholtz Center Berlin
Dr Werner is an X-Ray microscopy expert at HZB. Mammalian cells have a complicated structure with several cell organelles, each of which performs a specific cellular function. Filopodia are protrusions of the cell membrane that aid in cell migration.
He further stated, “For cryo X-Rray microscopy, the cell samples are first shock-frozen, so quickly that no ice crystals form inside the cell. This leaves the cells in an almost natural state and allows us to study the structural influence of external factors inside the cell.”
Study first author Michael Dyhr from Freie Universität Berlin stated, “Our work has already aroused considerable interest among experts.”
The neural network properly recognizes around 70% of the existing cell characteristics in a relatively short period, allowing for a very quick examination of the data set.
Dyhr concluded, “In the future, we could use this new analysis method to investigate how cells react to environmental influences such as nanoparticles, viruses, or carcinogens much faster and more reliably than before.”
Journal reference:
Dyhr, M. C. A., et al. (2023). 3D surface reconstruction of cellular cryo-soft X-ray microscopy tomograms using semisupervised deep learning. Proceedings of the National Academy of Sciences. doi.org/10.1073/pnas.2209938120