CRISPR/Cas is an efficient system for genetic engineering. It can precisely create a double-strand break at a target genomic location with only minimal modifications required to change the target location, thus bringing accuracy and versatility of mechanism, providing opportunities for the pharmaceutical industry within drug development.
It has improved disease modeling, identification of new drug targets, and may be suitable for therapeutic use.
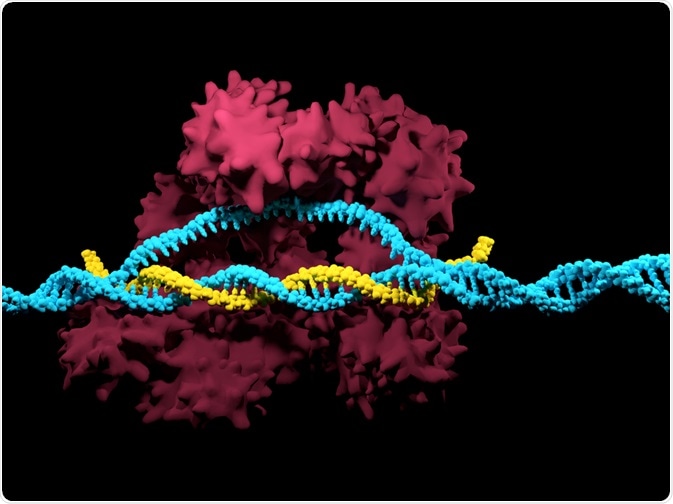
Image Credit: Meletios Verras/Shutterstock.com
CRISPR/Cas9 Overview
The CRISPR/Cas system is inherently simple, requiring only two components the guide RNA (gRNA) and a Cas (CRISPR associated) protein, normally Cas9.
The gRNA/Cas9 complex binds to the genome through the base pairing of the gRNA to DNA, while the Cas9 protein cuts the DNA backbone. Consequently, only the gRNA needs to be redesigned to adjust the targeting rather than the whole gRNA/Cas9 complex, thus providing a simpler and cheaper option.
CRISPR/Cas9 in disease model development
Due to its simplicity of only modifying the gRNA sequence, the CRISPR/Cas9 system speeds up the process of creating new models of disease.
Disease modeling has many advantages as it provides researchers the opportunity to study the disease in-depth, alongside studying the effect new drugs might have on the disease.
However, creating an accurate disease model for each disorder takes time and money. By both simplifying and speeding up the process, CRISPR/Cas9 facilitates the creation of additional disease models, thereby allowing drugs to be tested in the most accurate and suitable models.
The method for transfecting the cells with CRISPR depends on various factors including whether the editing is occurring in vivo or en vivo. Microinjection and electroporation are techniques which are suited to en vivo work.
They have been used to generate disease models from zygotes. However, this does carry the risk of mosaicism if the zygote divides before the genetic editing has been completed. In vivo techniques include the use of viruses to infect tissues with the CRISPR/Cas9 system.
Image Credit: Vladimir Staykov/Shutterstock.com
Adeno-associated viruses are one such method and have been used to generate animal models, but this technique is based on a naturally occurring virus, meaning the host organism may already carry antibodies against the delivery method.
Using CRISPR, models have been generated from a wide variety of organisms; from zebrafish to mice, pigs, monkeys, and even human cells. In human tissue, it has been used to create and cure cancer phenotypes. This demonstrates how many models can be made in different organisms using the CRISPR/Cas9 mechanism.
CRISPR/Cas9 as a drug target identifier
CRISPR can not only be used to speed up disease model creation, but it can also be used to subtly manipulate the proteome of the cell.
Many diseases are complex and arise from the interplay of many different genes, and therefore a refined method of manipulation is needed. Manipulation, where not only can the genes be knocked out or knocked in but also their level of expression can be controlled.
To this end, the Cas9 protein of the CRISPR/Cas system can be mutated, removing the endonuclease activity of domains that cut the DNA backbone, thus creating catalytically dead Cas9 (dCas9), a scaffold that can be precisely placed in the genome.
This dCas9 scaffold can be used to either promote gene transcription, CRISPRa or suppress it, CRISPRi. These techniques are used with large pools of different gRNAs in high throughput CRISPR library screening.
In CRISPR screens, large libraries of gRNAs are used that target different genomic locations. The phenotypes of all the cells are then studied to see which express the disease phenotype and consequently, which genes were targeted in that cell, and are therefore related to the disease.
These screens can either use CRISPR or CRISPRi, in loss of function screens, or CRISPRa, in a gain of function screens. Before CRISPR the only option for these large-scale screens was RNAi, which is less reliable than CRISPR, because RNAi may only lead to knockdown of genes not knockout.
However, there is one major drawback to CRISPR which is off-target effects. The gRNA/Cas9 complex might bind to multiple places within the genome which may lead to false identification of a disease-related gene.
Although there remain issues with the use of CRISPR, it is a very powerful tool for drug target identification. These CRISPR screens have resulted in the identification of numerous genes for drug target including cancer lethal genes involved in signaling, differentiation, survival, and regulatory processes.
CRISPR/Cas9 as a therapeutic.
As well as being used to search for new drugs, CRISPR can also be used as a therapeutic. One study involving lung cancer patients has reached the stage of a clinical trial.
T-cells were genetically edited with CRISPR/Cas9 to remove an immune checkpoint inhibitor which was identified as helping cancer cells avoid immune system detection. These genetically edited cells were then transfused back into the patient.
As with all genetic editing in human cells precautions must be taken to ensure there are no wanted side effects caused by off-target genetic editing.
Summary
CRISPR has been used in many aspects of the search for new drugs. By making disease models quicker and cheaper to produce, it has allowed more accurate models to develop. Within the cell, manipulation of the proteome has identified new drug targets that can be observed, and the effects of drugs studied.
Finally, there is the prospect of CRISPR being used directly as a therapeutic, however, careful measures must be put in place to limit unwanted side effects of off-target effects.

Image Credit: Syda Productions/Shutterstock.com
References
- Abrahimi, P. et al. (2015) ‘Efficient Gene Disruption in Cultured Primary Human Endothelial Cells by CRISPR/Cas9’, Circulation Research, 117(2), pp. 121–128. doi: 10.1161/CIRCRESAHA.117.306290.
- Ashmore-Harris, C. and Fruhwirth, G. O. (2020) ‘The clinical potential of gene editing as a tool to engineer cell-based therapeutics’, Clinical and Translational Medicine. Springer Berlin Heidelberg, 9(1). doi: 10.1186/s40169-020-0268-z.
- Broeders, M. et al. (2020) ‘Sharpening the Molecular Scissors: Advances in Gene-Editing Technology’, iScience. Elsevier Inc., 23(1), p. 100789. doi: 10.1016/j.isci.2019.100789.
- Cornet, C., Di Donato, V. and Terriente, J. (2018) ‘Combining Zebrafish and CRISPR/Cas9: Toward a more efficient drug discovery pipeline’, Frontiers in Pharmacology, 9, pp. 1–11. doi: 10.3389/fphar.2018.00703.
- Fellmann, C. et al. (2017) ‘Cornerstones of CRISPR-Cas in drug development and therapy’, Nature Reviews Drug Discovery, 16(2), pp. 89–100. doi: 10.1038/nrd.2016.238.
- Lino, C. A. et al. (2018) ‘Delivering CRISPR: A review of the challenges and approaches’, Drug Delivery. Informa Healthcare USA, Inc, 25(1), pp. 1234–1257. doi: 10.1080/10717544.2018.1474964.
- Luo, J. (2016) ‘CRISPR/Cas9: From Genome Engineering to Cancer Drug Discovery The CRISPR/Cas9 Endonuclease System’, Trends In Cancer, 2(6), pp. 313–324. doi: 10.1016/j.trecan.2016.05.001.
- Niu, Y. et al. (2014) ‘Generation of gene-modified cynomolgus monkey via Cas9/RNA-mediated gene targeting in one-cell embryos’, Cell, 156(4), pp. 836–843. doi: 10.1016/j.cell.2014.01.027.
- Ruan, J. et al. (2015) ‘Highly efficient CRISPR/Cas9-mediated transgene knockin at the H11 locus in pigs’, Scientific Reports. Nature Publishing Group, 5, pp. 1–10. doi: 10.1038/srep14253.
- Wanga, W. et al. (2016) ‘Delivery of Cas9 protein into mouse zygotes through a series of electroporation dramatically increased the efficiency of model creation’, Journal of Genetics and Genomics, 43(5), pp. 319–327. doi: 10.1016/j.jgg.2016.02.004.
Further Reading
Last Updated: May 12, 2020