Researchers from EMBL Barcelona’s Ebisuya group collaborated with colleagues from RIKEN, Kyoto University, and Meijo Hospital in Nagoya, Japan, to investigate oscillating patterns of gene expression, coordinated across time and space within a tissue grown in vitro.
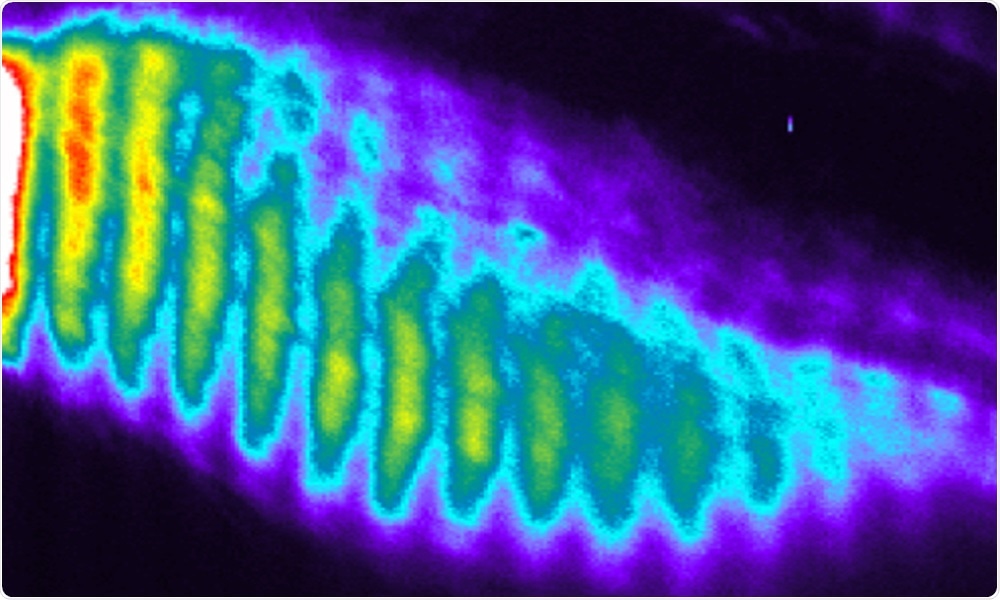
In this series of snapshots (kymograph), oscillations and waves of gene activity are seen in healthy cells. In cells with mutations in genes such as HES7, DLL3, and LFNG, these patterns of gene activity are stopped or disrupted, causing errors in development. Image Credit: Ebisuya group/EMBL.
The aim of the study is to explore the molecular causes of spondylocostal dysostosis (SCD)—a rare human hereditary disease. The study findings were published in Nature.
Segmentation clock
The vertebral column of humans is a highly repetitive structure with 33 vertebrae from top to bottom. This type of arrangement develops in the embryo by the orderly formation of a long row of structures named somites that later develop into the vertebrae and ribs.
This periodic arrangement of somites is produced by a cluster of genes called the segmentation clock. Molecular interactions within the cell lead to the oscillation of the expression of these genes, with a rise and fall in its genetic activity in a regular pattern over a period of time.
For every oscillation, one more somite is formed. Errors in this segmentation clock can lead to hereditary disorders of the vertebrae, like the rare condition SCD.
EMBL Research Scientist Mitsuhiro Matsuda and colleagues made efforts to develop a system to analyze the process in the laboratory since the dynamics of the human segmentation clock and associated diseases cannot be investigated directly in human embryos.
The researchers established cell lines that each lacked a gene considered as the causative mutation of SCD, which can be the reason for any of several genes, in different patients.
These cells were cultured to produce simplified versions of an embryo that possess several similar characteristic features. Cells that lacked a gene called HES7 did not exhibit oscillations, but cells that lacked the DLL3 and LFNG genes exhibited intact oscillations.
However, in spite of oscillations in these cell lines at the single-cell level, they did not coordinate properly across the tissue to create synchronized collective oscillations or traveling waves of gene activity.
Further tests
These experiments showed that the culture system developed by the researchers could demonstrate SCD mutations that had been designed into healthy cells. There arises a question of testing patients’ cells directly.
For this purpose, the scientists isolated a new cell line from a patient with a mutation in DLL3 and tested it in vitro. This cell line failed to exhibit traveling waves, as per predictions.
Researchers made use of the CRISPR–Cas9 gene-editing tool to rectify the patient’s mutation to provide a stronger piece of evidence that the DLL3 mutation was the reason behind it. As a result, the normal synchronization of the segmentation clock in the in vitro tissue was restored, thus corroborating that this specific mutation was responsible.
The segmentation clock, the mechanism underlying the periodic structures of the vertebral column, has been recapitulated in vitro. We also succeeded in evaluating two important properties of the segmentation clock separately: oscillation and synchronization. HES7, DLL3, and LFNG were already known as causative genes of SCD. But, for many SCD patients, the causative genes are still unknown. Our next goal is to identify a novel causative gene of SCD by using our newly established in vitro model.”
Miki Ebisuya, Group Leader, EMBL
Segmentation clock by EMBL scientists in the Ebisuya group
The oscillations and waves of the segmentation clock, seen in a culture of healthy human cells. Video Credit: Ebisuya group/EMBL.
Source:
Journal reference:
Matsuda, M., et al. (2020) Recapitulating the human segmentation clock with pluripotent stem cells. Nature. doi.org/10.1038/s41586-020-2144-9.