The genomic DNA found in eukaryotic cells is hierarchically wrapped by histones into chromatin. The structure and plasticity of chromatin play a significant role in the silencing and activation of gene transcription, and establish the fate of the cells during development and differentiation.
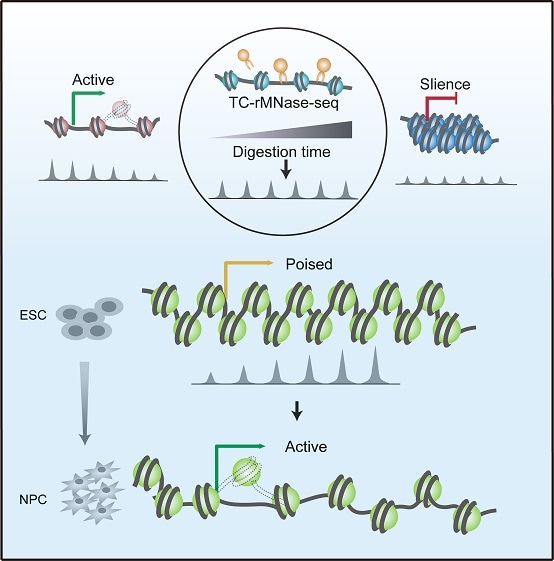
Model for the prediction of gene expression potential through identifying the local chromatin states. Image Credit: Dr LI Guohong’s group.
It is therefore a significant problem for the scientific frontier to analyze the status of chromatin and the transcription potentials of crucial development genes during the differentiation of cells.
Based on chromatin binding proteins and epigenetic markers, the chromatin states are split into several different types. The simultaneous presence of a couple of opposing chromatin modifications—repressive trimethylation at histone 3 lysine 27 (H3K27me3) and active trimethylation of histone3 lysine 4 (H3K4me3)—defines the poised state.
The poised state strongly associates with pluripotency in ESCs, with the ability to rapidly activate genes at the differentiation programs. But unlike epigenetic changes, the structural trait of the poised chromatin is still unclear and so does its association with gene expression potential.
In a research published in the Cell Reports journal, a team of researchers headed by Professor LI Guohong from the Institute of Biophysics (IBP) of the Chinese Academy of Sciences (CAS) and Professor XU Xueqing from Baoan Maternal and Child Health Hospital of Jinan University unraveled the structural traits of poised genes and offered a robust tool to estimate the transcription potential of genes.
Using a mild time-course MNase-seq technique (TC-rMNase-seq), the scientists discovered a pair of distinct groups of MNase-sensitive chromatin status in mouse embryonic stem (mES) cells, that is, “active” (Region I) as well as “poised” (Region II) chromatin regions, which exhibit opposing MNase-seq trends at the time of the digestion time course.
Based on this, the researchers created a computational technique to measure gene expression potential and chromatin status.
The scientists used a parameter Chromatin Opening Potential Index (COPI) and effectively scored the transcriptional potentials of the associated genes in mES cells in accordance with the different accessibilities at their promoters.
To analyze the utility of the technique, the researchers validated that genes having higher COPIs, like Ildr2, Scrt2, Ypel1, and Lfng, are poised in mES cells, but their promoters are partly opened and are instantly stimulated during the differentiation neuron progenitor cells (NPCs). Among them, Scrt2 and Lfng are known to work in NPCs and morphogenesis, in that order.
More significantly, through COPI prediction, the scientists observed that a pair of novel factors, Ypel1 and Ildr2, also play crucial roles during the differentiation of mouse NPCs in vitro.
The researchers proposed that COPI scores offer a robust tool to estimate and detect significant novel genes that play a role in a wide range of biological processes, such as key regulators during differentiation and development, and intermediate early genes in immune and neural responses.
Source:
Journal reference:
Yu, J., et al. (2020) Analysis of Local Chromatin States Reveals Gene Transcription Potential during Mouse Neural Progenitor Cell Differentiation. Cell Reports. doi.org/10.1016/j.celrep.2020.107953.