A team led by Karolinska Institute used artificial intelligence (AI) and structural biology to learn more about two similar proteins that protect against bacterial infection in the urinary tract and the gastrointestinal system. Their findings were published in the journal Nature Structural & Molecular Biology.
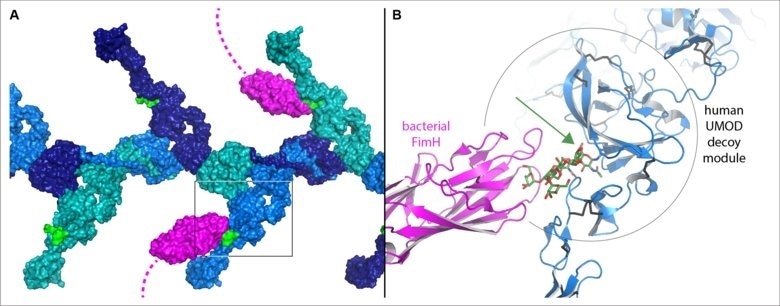
Figure 1. Model of the human UMOD/FimH interaction, assembled by combining the structural information described in the study. A, The high-mannose sugar chain (green) of the decoy modules branching out from a filament of UMOD (shades of blue) is bound by the FimH sugar-binding domain at the tip of bacterial adhesive filaments (magenta). B, Detail of the region boxed in (A), showing how the sugar chain recognized by FimH protrudes from the crevice between the two parts of the UMOD decoy module (green arrow). Image Credit: Luca Jovine
These findings could in principle be explored to develop a decoy alternative to conventional antibiotic treatments. And our structural work explains many human mutations associated with kidney diseases, as well as provides information on a protein region whose interaction with filtered light chains is implicated in cast nephropathy.”
Luca Jovine, Study Corresponding Author and Professor, Department of Biosciences and Nutrition, Karolinska Institute
Uromodulin (UMOD) is a protein whose filaments play a vital role in avoiding urinary tract infection, which is the most prevalent type of non-epidemic bacterial infection, affecting approximately 150 million people each year. It has a unique sugar chain that has not been modified like others and is still of the high-mannose type.
This acts as a decoy by imitating high-mannose sugar cords on the urinary tract’s surface, to which bacteria would generally attach via a sugar-binding protein called FimH at the edge of their hair-like appendages. Instead, the bacteria attach to the UMOD sugar chain and are eventually eliminated through the urine.
In recent times, evidence has accumulated indicating that glycoprotein 2 (GP2), a UMOD-like molecule secreted by the pancreas and intestine, holds a similar role in combating bacterial infections in the gastrointestinal system.
We wanted to determine which sugar chain of GP2 corresponded to the high-mannose chain of UMOD and understand how the proteins manage to keep these sugars ‘special’ so that they act as bait for bacteria.”
Luca Jovine, Study Corresponding Author, Professor, Department of Biosciences and Nutrition, Karolinska Institute
AI positioning itself as a tool in structural biology
The researchers obtained X-ray crystallographic and cryo-electron microscopy (EM) data on GP2 and UMOD in partnership with Daniele de Sanctis at the ESRF synchrotron in Grenoble and Marta Carroni at SciLifeLab in Stockholm.
However, its interpretation is complex by a number of technical problems. At this point, the researchers began working with DeepMind and the team behind AlphaFold—a machine learning model that drew a lot of attention from the structural biology community due to its efficiency in the 14th Critical Assessment of Structural Prediction competition.
Despite the fact that AlphaFold had never “seen” UMOD or GP2, its forecasts were extremely accurate. This enabled the team, which also included Bin Wu from Nanyang Technological University in Singapore and Nao Yamakawa from Lille University, to decipher the GP2 X-ray data and match low resolution cryo-EM maps of UMOD.
The research indicates that the functionally important decoy module of UMOD and GP2 is divided into two parts distinguished by a crevice. Although the sugar chains that connect bacteria are attached to different regions of the GP2 and UMOD sequences, the 3D structures demonstrate that their bases are located within the crevice in both instances.
This describes how these specific sugar chains are protected from alteration and can thus be used as hooks to catch pathogenic bacteria.
Source:
Journal reference:
Stsiapanava, A., et al. (2022) Structure of the decoy module of human glycoprotein 2 and uromodulin and its interaction with bacterial adhesin FimH. Nature Structural & Molecular Biology. doi.org/10.1038/s41594-022-00729-3.