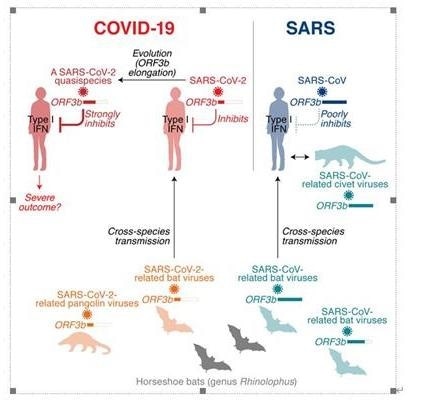
One of the features distinguishing SARS-CoV-2 from its more pathogenic counterpart SARS-CoV is the presence of premature stop codons in its ORF3b gene. Here, we show that SARS-CoV-2 ORF3b is a potent interferon antagonist, suppressing the induction of type I interferon more efficiently than its SARS-CoV ortholog. Phylogenetic analyses and functional assays reveal that SARS-CoV-2-related viruses from bats and pangolins also encode truncated ORF3b gene products with strong anti-interferon activity. Furthermore, analyses of approximately 17,000 SARS-CoV-2 sequences identify a natural variant in which a longer ORF3b reading frame was reconstituted. This variant was isolated from two patients with severe disease and further increased the ability of ORF3b to suppress interferon induction. Thus, our findings not only help to explain the poor interferon response in COVID-19 patients but also describe the emergence of natural SARS-CoV-2 quasispecies with an extended ORF3b gene that may potentially affect COVID-19 pathogenesis. Image Credit: ©Kei Sato.
One significant characteristic that differentiates COVID-19 from SARS in terms of immune reactions is the poor induction of a type I interferon (IFN) reaction by SARS-CoV-2 when compared to influenza A virus and SARS-CoV.
Impaired IFN responses are particularly linked to COVID-19 disease. But in in SARS-CoV-2 infection, the molecular mechanisms underlying the inefficient IFN reactions continue to remain unclear.
A team of researchers from The Institute of Medical Science, The University of Tokyo (IMSUT) aimed to define the viral factor(s) that determine immune activation following SARS-CoV-2 infection and discovered that the ORF3b gene, encoded by SARS-CoV-2, is a powerful IFN antagonist.
The poor IFN responses in COVID-19 patients may be explained by the action of this viral product, ORF3b.”
Kei Sato, Lead Scientist and Associate Professor (Principal Investigator), Division of Systems Virology, Department of Infectious Disease Control, The Institute of Medical Science, The University of Tokyo
The study results were published in the Cell Reports journal on September 4th, 2020.
ORF3b as a viral IFN antagonist
Even though SARS-CoV infection leads to severe and acute and pneumonia, it may be asymptomatic or lead to flu-like symptoms like cough, fever, and fatigue. Moreover, COVID-19 is the poor induction of a type I interferon (IFN) when compared to SARS-CoV and influenza A virus infections, a characteristic of SARS-CoV-2 infection.
As stated above, impaired IFN responses are specifically linked to the severity of COVID-19 disease. But in SARS-CoV-2 infection, the molecular mechanisms underlying the inefficient IFN responses continue to be vague.
When the team compared the sequences of SARS-CoV-2-encoding genes to those of SARS-CoV, they found that the length of the SARS-CoV-2 ORF3b gene is distinctly shorter than that of SARS-CoV ORF3b.
Since SARS-CoV ORF3b is recognized as a viral antagonist against the production of IFN, the researchers hypothesized that the variation in the length of the ORF3b gene between SARS-CoV and SARS-CoV-2 may modify their anti-IFN activity and may further describe the variation in the symptoms of both these viral infections.
Unexpectedly, SARS-CoV-2 ORF3b is a more powerful IFN antagonist when compared to SARS-CoV ORF3b. Functional assays and phylogenetic analyses showed that SARS-CoV-2-related viruses from pangolins and bats also encode shorter products ofORF3b genes with potent anti-IFN activity.
Characterization of a natural SARS-CoV-2 ORF3b variant with enhanced anti-IFN activity
Moreover, analyses of around 17,000 SARS-CoV-2 sequences detected a natural variant, wherein a longer ORF3b reading frame was reconstituted. This natural variant inhibits IFN much more efficiently when compared to ORF3b of the parental strain of SARS-CoV-2.
In line with an association of IFN suppression with the severity of disease, the two patients in Ecuador, who were harboring SARS-CoV-2 with the extended ORF3b variant, were severely ill—one died of COVID-19 and the other was treated in an intensive care unit.
But most significantly, there is no direct proof to indicate that the viruses identified in both these COVID-19 patients in Ecuador are more pathogenic than the reference strain. Even though it was impossible to differentiate whether this variant is linked to a different result in disease, it is viable that naturally occurring length variants of ORF3b can possibly contribute to the emergence of SARS-CoV-2 variants that are more several pathogenic.
Therefore, viral sequences should be continuously supervised to observe whether new ORF3b variants emerge during the present pandemic.
To our knowledge, this study is the first investigation revealing the role of a SARS-CoV-2-encoded protein that can be associated with the progression of COVID-19.”
Kei Sato, Lead Scientist and Associate Professor (Principal Investigator), Division of Systems Virology, Department of Infectious Disease Control, The Institute of Medical Science, The University of Tokyo
Source:
Journal reference:
Konno, Y., et al. (2020) ISARS-CoV-2 ORF3b Is a Potent Interferon Antagonist Whose Activity Is Increased by a Naturally Occurring Elongation Variant. Cell Reports. doi.org/10.1016/j.celrep.2020.108185.