Kanazawa University researchers have reported a new approach of atomic force microscopy for imaging biological processes and specimens. The technique provides higher frame rates and causes less disruption to samples. The study has been reported in the Review of Scientific Instruments.
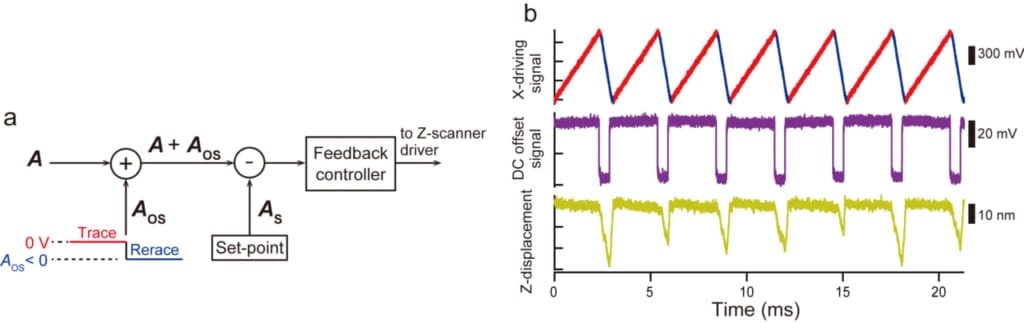
The circuit installed for the OTI mode and its operation. (a) During retrace scanning, a DC offset signal (Aos < 0) is added to the amplitude signal (A). The feedback control operates as if the probe were in strong contact with the sample, and thus the sample stage is moved away from the tip. (b) Driving signal for X-scanner in the OTI mode (top), DC offset signal added to true amplitude signal (middle), and Z-scanner displacement (bottom). Image Credit: Kanazawa University.
High-speed atomic force microscopy (HS-AFM) is an imaging method used to visualize biological processes such as protein activity. Today, standard HS-AFM frame rates are as high as12 frames per second; however, better video rates are needed to enhance the capabilities of the technique so that it can be extended to an ever-growing range of biological specimens.
Furthermore, faster recording times mean less interaction between the specimen and the probe—a tip scanning the surface of the sample—rendering the imaging process less invasive.
Shingo Fukuda and Toshio Ando from Kanazawa University’s Nano Life Science Institute (WPI-NanoLSI) have now created an alternative HS-AFM method to boost the frame rate by up to 30 frames per second.
An AFM image is created by laterally shifting a tip just above the surface of a sample. During this XY-scanning movement, the position of the tip in the direction perpendicular to the XY-plane (the z-coordinate) will trace the height profile of the sample. The difference in the z-coordinate of the tip creates a heightmap—the image of the specimen.
Fukuda and Ando also investigated the HS-AFM technique in the supposed amplitude-modulation mode. The tip is subsequently made to oscillate at a fixed amplitude. Due to height variations in the structure of the sample, the oscillation amplitude will change when scanning a surface.
To return to the original amplitude, the tip-sample distance must be corrected. The extent of the correction to be made is determined by the surface topology of the sample and is governed by the supposed feedback control error of the setup.
The researchers discovered that this feedback control error differs when the tip travels in reverse directions, a process known as tracing and retracing. This disparity is eventually caused by the varied physical forces at play when the tip is “pulled” (tracing) and when it is “pushed” (retracing).
Based on their understanding of the physics of the tracing and retracing procedure, both Fukuda and Ando created an imaging regime that avoids retracing. This should be properly considered in the control algorithm.
The only-trace-imaging mode was tested on actin filament samples (actin is a protein commonly found in cells.) Not only was the imaging quicker, but it was also less invasive—the filaments did not beak much frequently. The researchers also recorded polymerization processes (via protein-protein interactions); once again, the technique was found to be less disruptive and faster than the typical AFM tracing-retracing operations.
The researchers are optimistic that their “simple and highly effective method will soon be installed in the existing and upcoming HS-AFM systems and will improve a wide range of HS-AFM imaging studies in biophysics and other fields.”
Source:
Journal reference:
Fukuda, S & Ando, T (2021) Faster high-speed atomic force microscopy for imaging of biomolecular processes. Review of Scientific Instruments. doi.org/10.1063/5.0032948.