Researchers from the Agency for Science, Technology, and Research (A*STAR)’s Institute of Molecular and Cell Biology (IMCB) recently constructed the first single-cell atlas of the human and porcine eyes. This enabled them to generate a disease map of genes connected with eye disorders across various cell types, and also as the major switches which regulate cell specialization of individual ocular tissues.
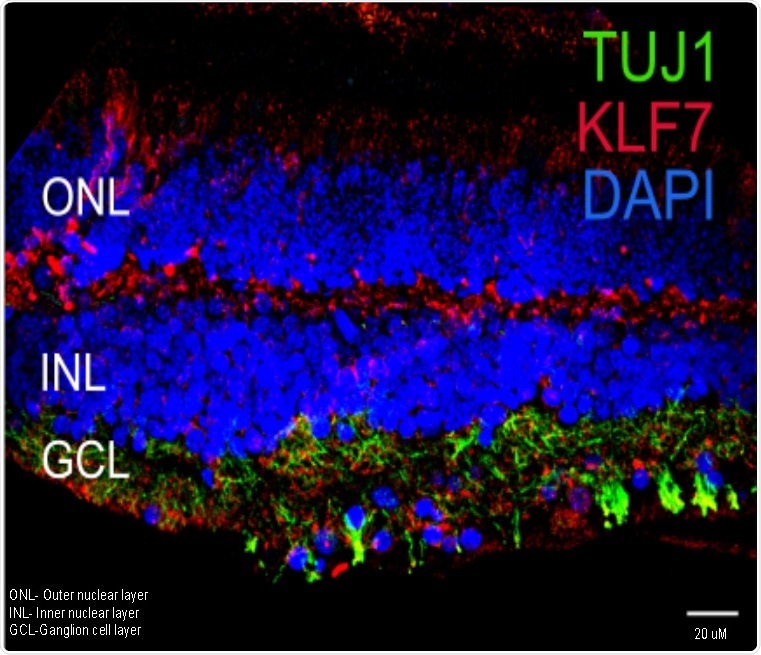
KLF7(red), which could accelerate retinal ganglion cell (RGC) generation, is generally expressed in the RGC layer marked by TUJ1+ neurons (green) of retina. Lab grown RGCs could replace damaged RGCs as a potential therapy for eye disorders. Image Credit: Pradeep Gautam, Institute of Molecular and Cell Biology (IMCB), A*STAR.
The study could offer a novel understanding of human eye diseases and age-related eye disorders. This could lead the light for creating cell replacement therapies and regenerative medicine for eye diseases. The study was published on September 28th, 2021in the leading scientific journal Nature Communications.
The group of scientists headed by IMCB, included investigators from the National University of Singapore (NUS), Nanyang Technological University (NTU), Singapore Eye Research Institute (SERI), Mayo Clinic, Icahn School of Medicine at Mount Sinai, University of Melbourne, and Shenzhen Eye Hospital.
The human eye has various cell types from different functional roles and developmental origins. To decode this diversity of cellular functions employing single-cell RNA sequencing, the scientists cataloged more than 50,000 cells in porcine and human eyes and created a cell atlas of these eyes that differentiates individual cells by their gene activity.
Some eye disorders are linked to mutations in the human genes, hence the atlas which profiles disease-causing genes throughout all cell types of the eye can provide a better knowledge of how cell types are impacted by such disorders.
A map of cell surface proteins that perform as viral entry receptors in the human eye can provide insight into how infections travel via the ocular route. The study demonstrated that TMPRSS2 and ACE2, the primary cell surface proteins responsible for the entry of SARS-CoV-2 into the human, are expressed in the eye’s conjunctival cells. Other studies too reported similar findings indicating that an infection from the ocular route is likely.
It is fascinating to observe the cross-talk of ocular cells through the interaction of signaling molecules with receptors. This will help us to understand how individual cell responds to external factors like injury.”
Pradeep Gautam PhD, Study Co-First Author and Scholar, Institute of Molecular and Cell Biology, Agency for Science, Technology, and Research
Eye diseases like glaucoma—the major reason for blindness—occurs due to the degeneration of retinal ganglion cells (RGCs). Employing embryonic stem cells as a platform for differentiation, the scientists developed RGC progenitor cells in culture and evaluated that a switch KLF7 could speed up RGC generation, unraveling opportunities for the cultivation of laboratory-grown RGCs to substitute damaged RGCs as a potential therapy.
This study paves the way to create mature retinal ganglion cells in the lab, which could be harnessed for the development of new approaches to reverse vision loss that results from optic nerve degeneration, including glaucoma.”
Dr Jonathan Loh Yuin-Han, Research Director, Institute of Molecular and Cell Biology, Agency for Science, Technology, and Research
Dr Jonathan Loh Yuin-Han is the lead researcher of the study.
The researchers intend to employ RGCs for cell therapies in patients and are investigating further to evaluate the KLF7-derived RGCs for preclinical researches. The knowledge of key molecular switches and their capability to engineer cell types in culture, laboratory-grown cells offer a key to therapy.
Source:
Journal reference:
Gautam, P., et al. (2021) Multi-species single-cell transcriptomic analysis of ocular compartment regulons. Nature Communications. doi.org/10.1038/s41467-021-25968-8.