Genome sequencing, in which researchers use laboratory approaches to predict the genetic makeup of a particular organism, is becoming more common in insect studies. A better comprehension of insect biology allows researchers to effectively manage insects that are advantageous to the ecosystem as well as those that harm the food supply and endanger human health by transporting diseases.
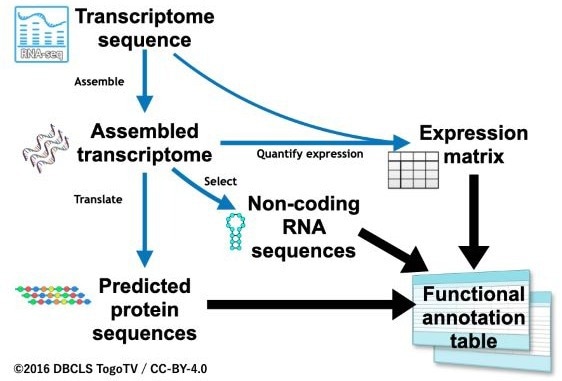 Overview of the functional annotation workflow, Fanflow4Insects. Fanflow4Insects consists of three pipelines. Image Credit: Hidemasa Bono licensed under CC-BY 4.0
Overview of the functional annotation workflow, Fanflow4Insects. Fanflow4Insects consists of three pipelines. Image Credit: Hidemasa Bono licensed under CC-BY 4.0
Fanflow4Insects is a workflow method developed by researchers that annotates gene functions in insects. Researchers use functional annotation to learn about the biological identity of a gene. The new technique developed by the team makes use of transcribed sequence data as well as genome and protein sequence datasets.
The team used Fanflow4Insects to jot down the useful details of the Japanese stick insect and silkworms, such as gene expression and sequence analysis. Their workflow’s functional annotation data will tremendously improve the accessibility of entomological studies using genome editing.
The technique, developed by researchers from Hiroshima University, Tokyo University of Agriculture and Technology, and the RIKEN Center for Integrative Medical Sciences, was published in the journal Insects on June 27th, 2022.
Since insects are so varied and abundant, researchers require a method to study them on a massive scale. This is what prompted researchers to start working on sequencing insect genomes. Scientists had decoded and registered the genomes of approximately 3000 insect species as of May 2022. They are also using long-read sequencing technology to streamline the sequencing of insect genomes.
Next-generation sequencing has made it much easier for scientists to decode the genomes and transcript sequences of multiple insects. The biological viewpoint of these sequences, however, remains a major bottleneck in transcriptome analysis.
The transcriptome is the total number of RNA molecules in an organism. Transcriptome analysis is a critical first stage in functional annotation, and it provides important information for selecting genome editing targets.
Since some insects’ genomes are larger than the human genome, the tough process of whole-genome sequencing is made even harder. As a result, researchers are using transcriptome sequencing in conjunction with next-generation sequencing technology, also known as RNA sequencing, to evaluate large genome-size insects.
By gathering tens of millions of reads, researchers can efficiently identify tens of thousands of potential genes in a particular tissue using this valuable tool. The gene sequences are then assembled into transcriptional units for recognition.
However, this type of analysis is dependent on researchers having access to huge datasets with functional annotation. Databases do appear, but they are unable to keep up with the increased sequencing of insect genomes.
As transcriptome analysis becomes more prevalent, many research teams are operating their pipelines, with data about transcription units from different studies reported research by research. These pipelines are collections of algorithms that are used to perform genome sequencing data.
However, researchers require a method to incorporate functional annotation from all of the different groups conducting this type of research into public databases.
The research group used their recently developed Fanflow4Insects to build a functioning annotation pipeline for the silkworm in this present study. The scientists then used Fanflow4Insects to analyze the transcriptomes of the Japanese stick insect.
Functional annotation is one of the most important processes to accelerate the selection of target genes once the genome or transcriptome of the target organism is decoded. The functional annotation information obtained by the workflow Fanflow4Insects will greatly expand the possibilities of entomological research using genome editing.”
Hidemasa Bono, Study First Corresponding Author and Professor, Graduate School of Integrated Sciences for Life, Hiroshima University
The Fanflow4Insects workflow for insects has been publicly established and is freely available on GitHub. The information from Fanflow4Insects can be used to compare insects with different phenotypes when combined with the functional annotation obtained from expression.
“Using Fanflow4Insects, we are going to annotate insects that produce useful substances. The ultimate goal of this study is to make it possible to design molecular networks in insects using computer simulation,” said Bono.
Source:
Journal reference:
Bono, H., et al. (2022) Systematic Functional Annotation Workflow for Insects. Insects. doi.org/10.3390/insects13070586.