Living cell factories can manufacture custom drug compounds and biofuels using biological enzymes. Researchers at the Chalmers University of Technology have created a computer model that can anticipate how quickly enzymes function. Courtesy of this model, it is now possible to locate the most effective living factories and study challenging diseases.
 The researchers tested their model by simulating metabolism in more than 300 types of yeasts. When compared with measured, pre-existing knowledge, the researchers concluded that models with predicted kcat values could accurately simulate metabolism. The image shows common baker’s yeast, Saccharomyces cerevisiae. Image Credit: Chalmers University of Technology
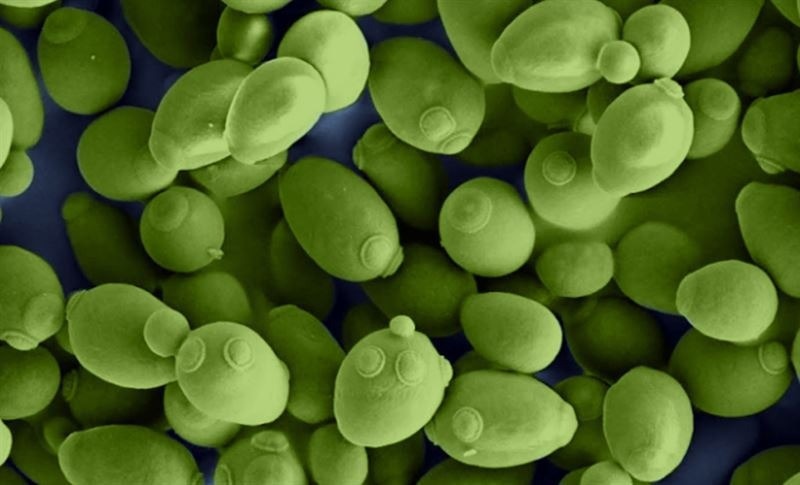
The researchers tested their model by simulating metabolism in more than 300 types of yeasts. When compared with measured, pre-existing knowledge, the researchers concluded that models with predicted kcat values could accurately simulate metabolism. The image shows common baker’s yeast, Saccharomyces cerevisiae. Image Credit: Chalmers University of Technology
All living cells contain proteins known as enzymes. Their function is to accelerate the rate of particular chemical reactions occurring in cells by acting as catalysts. Therefore, enzymes are like nature’s mini factories and are essential to the functioning of life on Earth.
Furthermore, they are employed in the production of pharmaceuticals, sweeteners, dyes, and detergents, among other products. Although there are countless possible applications, studying the enzymes is expensive and time-consuming, which limits the possibilities.
To study every natural enzyme with experiments in a laboratory would be impossible, they are simply too many. But with our algorithm, we can predict which enzymes are most promising just by looking at the sequence of amino acids they are made up of.”
Eduard Kerkhoven, Study Lead Author and Researcher, Systems Biology, Chalmers University of Technology
Only the most promising enzymes need to be tested
Understanding the metabolism of a cell requires an understanding of the enzyme turnover number, often known as the kcat value, which indicates how quickly and effectively an enzyme operates.
Researchers from Chalmers University have created a computer model in the current study that can swiftly determine the kcat value. The order of the amino acids that make up the enzyme is the only information required, and this information is frequently easily accessible in open databases.
Only the enzymes that seem most likely to succeed must undergo laboratory testing once the model makes its initial selection.
The researchers think that the new calculating model could be quite significant given the abundance of naturally occurring enzymes.
“We see many possible biotechnological applications. As an example, biofuels can be produced when enzymes break down biomass in a sustainable manufacturing process. The algorithm can also be used to study diseases in the metabolism, where mutations can lead to defects in how enzymes in the human body work,” stated Kerkhoven.
More knowledge on enzyme production
Production of goods using natural organisms is more efficient than using industrial procedures, which opens up more potential uses. Among these is the antibiotic penicillin taken from a mold, the cancer medication taxol from yew, and the sweetener stevia. Normally, natural organisms create these in little quantities.
Kerkhoven added, “The development and manufacture of new natural products can be greatly helped by knowledge of which enzymes can be used.”
The calculating model can also highlight variations in kcat value brought on by enzyme mutations and point out unintended amino acids that could significantly reduce an enzyme’s effectiveness. If the enzymes produce multiple “products,” the model can also predict this.
“We can reveal if the enzymes have any ‘moonlighting’ activities and produce metabolites that are not desirable. It is useful in industries where you often want to manufacture a single pure product,” further stated Kerkhoven.
By simulating the metabolism of more than 300 different species of yeast using 3 million kcat values, the researchers tested their model. They developed computer simulations of how rapidly the yeasts could reproduce themselves or make particular byproducts, like ethanol.
The researchers concluded that models with projected kcat values could accurately replicate metabolism when compared to measured, prior information.
Source:
Journal reference:
Li, F., et al (2022) Deep learning-based kcat prediction enables improved enzyme-constrained model reconstruction. Nature Catalysis. doi:10.1038/s41929-022-00798-z