Researchers from the National University of Singapore (NUS) have shown that, compared to conventional evaluation methods, deep learning enables them to detect the dynamics of single molecules more precisely and with fewer data.
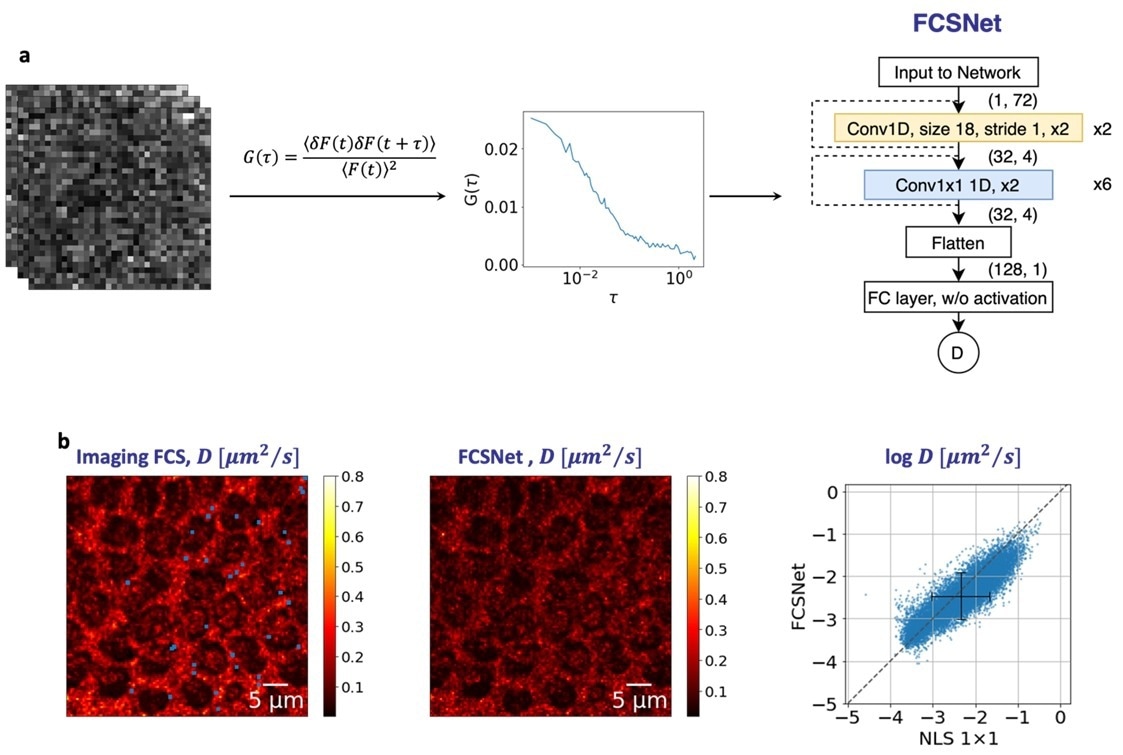 Figure. (a) FCSNet is a convolutional neural network (CNN), which takes in the autocorrelation function of a measurement as an input to predict the diffusion coefficient. (b) FCSNet allows evaluations in complex samples with a precision as good, or better than traditional Imaging FCS methods, as shown by the heatmaps. Image Credit: Biophysical Journal
Figure. (a) FCSNet is a convolutional neural network (CNN), which takes in the autocorrelation function of a measurement as an input to predict the diffusion coefficient. (b) FCSNet allows evaluations in complex samples with a precision as good, or better than traditional Imaging FCS methods, as shown by the heatmaps. Image Credit: Biophysical Journal
Convolutional neural networks (CNNs) were employed to study the motion of individual molecules in cells, artificial systems, and small animals. This technique should speed up single-molecule measurements in intricate systems and increase accessibility for a larger group of scientists.
The smallest unit of biological systems that may be observed is a single molecule. By comprehending its behavior and interactions, users can gain insights into how biological systems function, which opens the door to targeted therapies for disorders. Fluorescence spectroscopy is one of the most effective methods for observing single molecules.
This is because of its specificity, which enables the observation of only labeled molecules and powerful signal. Fluorescence Correlation Spectroscopy (FCS) has been employed in this sector for more than 50 years, providing very accurate and precise measurements of molecular mobility and interaction.
Imaging FCS, a more recent development of this method, expands its potential to characterize mobilities, concentrations, and interactions among other parameters in entire images. Imaging FCS is capable, but it has limitations because it needs a lot of data (about 100 MB for each measurement). Evaluations are sluggish as a result of the high computational processing required for this.
Under the direction of Professor Thorsten Wohland from the Department of Biological Sciences and Chemistry and Adrian Röllin from the Department of Statistics and Data Science at NUS, a research team used deep learning techniques to achieve results that were comparable to traditional methods while requiring less data roughly 5 MB for each measurement.
The study team members Dr Wai Hon Tang and Mr. Shao Ren Sim developed two CNNs, FCSNet and ImFCSNet, which are used in this technique. CNNs are a class of deep learning algorithms that are well-suited for visual data analysis. They use several layers of specialized filters to search the image for particular elements like edges, textures, and colors.
They get a deeper comprehension of the image by gradually extracting and integrating these qualities, which enables them to identify patterns and objects within the visual input.
In terms of diffusion coefficients, both the FCSNet and ImFCSNet-based approaches are more accurate than conventional FCS methods; however, they differ in the trade-offs they make about data requirements and spatial resolution. These methods have the potential to significantly reduce the evaluation time by using less data, especially for huge datasets or complicated systems.
The Biophysical Journal published its findings.
These CNNs are trained on simulated data and can predict diffusion coefficients from much smaller datasets compared to traditional FCS methods. They are also model agnostic and can be used with any microscope setup.”
Thorsten Wohland, Professor, Department of Biological Sciences and Chemistry, National University of Singapore
Dr. Tang added, “CNNs revolutionize data analysis, offering accelerated and simplified evaluation processes. While it is important to validate their performance, CNNs have the potential to make powerful techniques available to the wider research community.”
The group anticipates that this method will create new avenues for accelerating single-molecule research and increasing the technology’s accessibility for a larger user base. The CNNs are ready to democratize FCS and do not need professional input.
Source:
Journal references:
Wai Hoh Tang, et.al., (2024). Deep learning reduces data requirements and allows real-time measurements in Imaging FCS. Biophysical Journal. https://doi.org/10.1016/j.bpj.2023.11.3403
Krieger, J. W. et.al., (2024). Imaging fluorescence (cross-) correlation spectroscopy in live cells and organisms. Nature Protocols. https://doi.org/10.1038/nprot.2015.100