Metagenomics refers to the study of genetic material from an environmental specimen. This field is now progressing as organisms evolve or are identified worldwide.
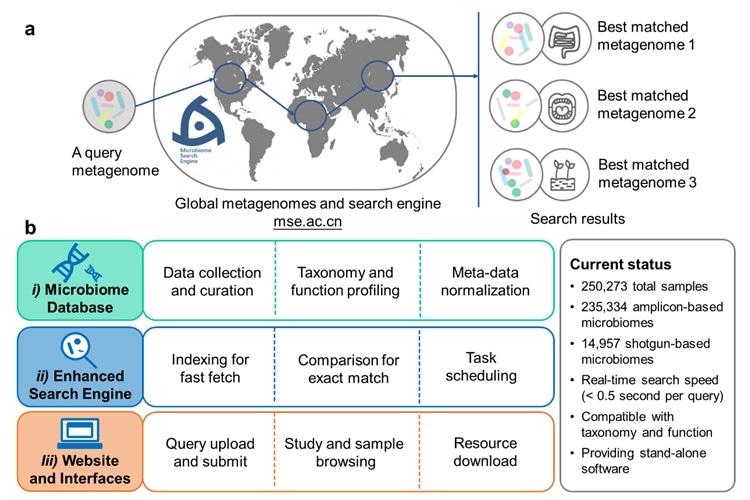
Design of the Microbiome Search Engine 2 (MSE 2). Image Credit: Jing Gongchao.
To associate the recently designed microbiomes with currently available data sets, a research team based in China has designed a novel Microbiome Search Engine 2 (MSE 2). The study was published in the American Society for Microbiology journal, mSystems, on January 19th, 2021.
Here, we introduce MSE 2, a microbiome database platform for searching query microbiomes in the global metagenome data space based on taxonomic or functional similarity of the whole microbiome.”
Jing Gongchao, Study Co-First Author, Single-Cell Center, CAS Key Laboratory of Biofuels, Qingdao Institute of BioEnergy and Bioprocess Technology, Chinese Academy of Sciences
The prior model of the search engine restricted the questions relating to taxonomic similarities, which means the functional genes should match. There was no other way for scientists to compare different types of specimens that carried out the same role in different microbiomes.
According to JING, the potential to explore how the same kind of function could have emerged in diverse bacteria may provide guidance for detecting and treating various diseases.
A search-based strategy is useful for large-scale mining of microbiome datasets, providing a bird's eye view of the microbiome data space and disease diagnosis via microbiome big data. The new ability to search the microbiome space via functional similarity greatly expands the scope of search-based mining of the microbiome big data.”
Jing Gongchao, Study Co-First Author, Single-Cell Center, CAS Key Laboratory of Biofuels, Qingdao Institute of BioEnergy and Bioprocess Technology (QIBEBT), Chinese Academy of Sciences
“By adding a function-based dimension for these and related applications, MSE 2 should accelerate large-scale mining of the ever-expanding metagenome data space,” added Jing.
MSE 2 features an expanded database with meta-data obtained from a total of 819 research works, that is, updated data compatibility to better integrate an easy-to-use interface and the recently available data sets. A single question takes less than half a second to look for the whole database of over 260,000 specimens.
Over the past decade we have been passionately collecting published microbiome data – they record the kinds of microbe species and the types of microbial communities that have ever lived on our planet. By collecting and curating them in a minable database, MSE 2 allows these invisible but pivotal creatures on Earth to be "remembered" by future generations, and those scientists who first discovered them to be recognized.”
LIU Lu, Study Co-First Author, Single-Cell Center, CAS Key Laboratory of Biofuels, Qingdao Institute of BioEnergy and Bioprocess Technology, Chinese Academy of Sciences
Source:
Journal reference:
Jing, G., et al. (2021) Microbiome Search Engine 2: a Platform for Taxonomic and Functional Search of Global Microbiomes on the Whole-Microbiome Level. mSystems. doi.org/10.1128/mSystems.00943-20.