Legumes, or plants belonging to the bean family, form nodules on their roots to absorb nitrogen. These plants tend to stop the production of nodules when nitrogen is abundant. However, it has been a mystery how exactly the occurrence of nitrate regulates nodule formation in these plants.
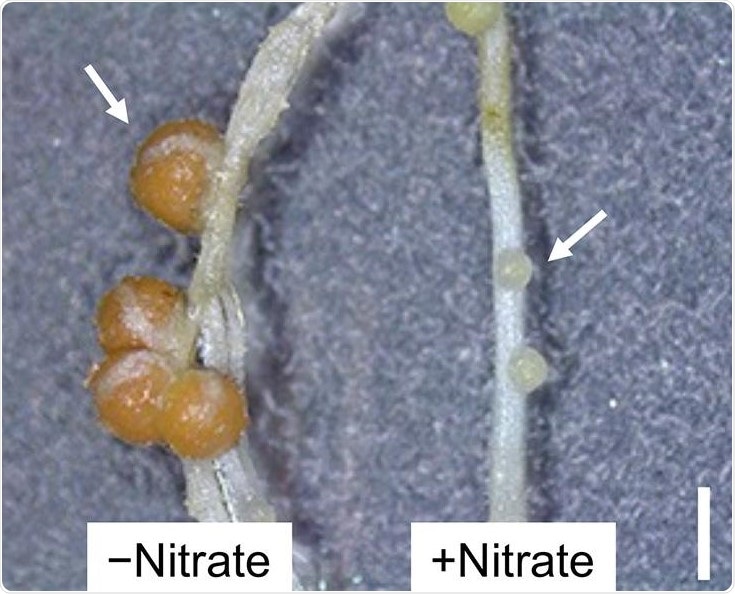
Nitrate-induced control of root nodule formation. Legumes halt root nodule formation when plentiful amounts of nitrogen nutrients, such as nitrate, are present in the environment. Arrows indicate root nodule. Scale bar: 1 mm. Image Credit: University of Tsukuba.
Scientists from Japan have now discovered that interactions between nitrate and proteins can stimulate and repress genes, thereby regulating nodulation with prospective applications in sustainable agriculture.
A team of scientists from the University of Tsukuba has demonstrated that the different DNA-binding characteristics between proteins initiate nodule development govern whether genes that take part in a symbiosis that regulates nodulation switch on or off and whether this gene expression is induced by the presence of nitrate. The study was recently published in The Plant Cell journal.
So far, insights into the molecular activity that governs how legumes halt nodulation in the presence of excess nitrate were incomplete. Earlier studies found that transcription factors, or proteins that help switch particular genes “on” or “off,” contributed to nodule formation. However, that is only a part of the theory.
Building on the previous identification of transcription factors for proteins (known as NLPs) involved in nodule inception, we sought to answer the question of how symbiotic gene expression facilitating nodulation is controlled by nitrate. We tested specific NLPs and found that they have overlapping functions, causing nitrate-induced control of nodulation.”
Takuya Suzaki, Study Senior Author and Professor, University of Tsukuba
The researchers investigated these molecular interactions by studying RNA molecules and plant traits with the help of proteins from Lotus japonicus. Certain proteins were found to exhibit dual functions, serving as master regulators for nitrate-dependent gene expression.
The team also found new protein binding sites and compared them with previously identified ones. The study results unravel fundamental principles associated with NLP-regulated transcription of symbiotic genes that inhibit nitrate nodulation.
The researchers highlighted further questions. Certain NLPs occur in cell nuclei in response to nitrate and inhibit the production of nodules. Some other NLPs continuously aggregate in nuclei regardless of the nitrate levels. In the case of the latter, it is not clear how they work specifically when nitrate is present.
The location of the NLPs in the cell is highly crucial because translation (when RNA is coded into proteins) occurs in the cytoplasm of the cell. If changes to proteins take place following the reading of the genetic code (post-translational modifications), it could account for how these NLPs access protein-protein interactions and control genes.
Uncovering how transcription factors influence gene expression has been a missing piece to the puzzle of understanding plant transcription regulation. Our discoveries bring us closer to knowing what is possible within these complex molecular relationships, but there is plenty left to untangle. Future research should aim to answer the question of how nodulation is regulated by other NLPs and in other plant species of interest.”
Takuya Suzaki, Study Senior Author and Professor, University of Tsukuba
Source:
Journal reference:
Nishida, H., et al. (2021) Different DNA-binding specificities of NLP and NIN transcription factors underlie nitrate-induced control of root nodulation. The Plant Cell. doi.org/10.1093/plcell/koab103.