Soil carbon has been an important topic of study in climate change due to the increasing prominence of greenhouse effects. The primary agents for the transformation of soil carbon are soil microbes.
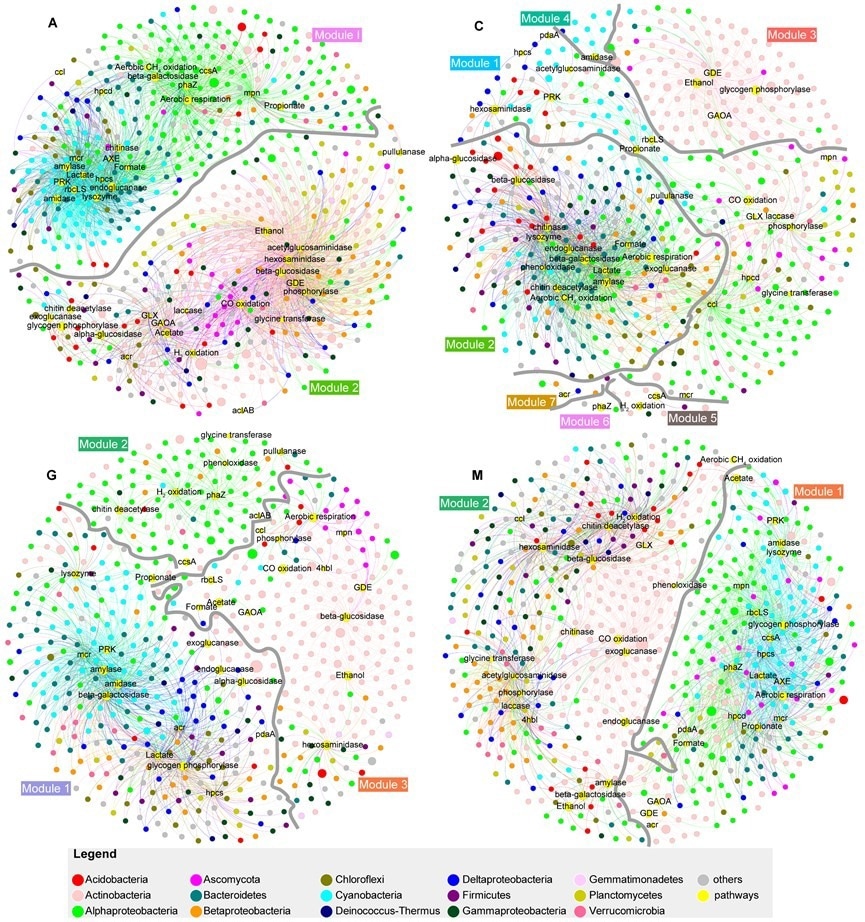 Co-occurrence networks for genes related to carbon cycle and high-abundance genera for each biocrust successional stage. Image Credit: Institute of Hydrobiology
Co-occurrence networks for genes related to carbon cycle and high-abundance genera for each biocrust successional stage. Image Credit: Institute of Hydrobiology
Metagenomics can make it challenging to understand the pattern and process of the soil carbon cycle at the level of the microbial community due to the complexity of elements such as microbial physiology, the content of organic compounds in soils, and diversity among redox forms.
Biocrusts are perfect for modeling since they are prevalent in arid areas, make up more than 40% of the world’s land surface, and are dominated by cryptophytes like Cyanobacteria, lichens, and mosses for research of soil ecosystems employing metagenomics. Additionally, they can stop sandy soils from being eroded by wind, fostering the growth and succession of ecosystems.
Researchers from the Institute of Hydrobiology (IHB) of the Chinese Academy of Sciences, under the direction of Professor Chunxiang Hu, examined successional differences in biocrusts as well as the relationships between C-cycle processes, environmental factors, and microorganism groups in a study that was published in Soil Biology and Biochemistry journal.
The scientists used four different types of biocrusts to separate various successional stages, namely cyanobacteria-dominated crusts (A), cyanolichen-dominated crusts (C), chlorolichen-dominated crusts (G), and moss-dominated crusts (M). In the last four years, they have been collected numerous times from Shapotou District in the Tengger Desert.
Using metagenomic sequencing, the scientists found a low abundance of genes associated with light-driven inorganic carbon fixation and a high abundance of genes associated with chemical energy-driven degradation of macromolecular organic carbon (OC), fermentation, aerobic respiration, and CO oxidation.
For OC decomposition, we found that genes mediating starch/glycogen and cellulose degradation were most abundant during the initial complex OC degradation, as were genes mediating fermentation during terminal steps of OC decomposition.”
Chunxiang Hu, Professor, Institute of Hydrobiology, Chinese Academy of Sciences
The researchers applied the metagenomic information with absolute quantification through the use of GeoChip and key enzyme activity measurements to evaluate successional changes in the carbon cycle.
We observed that inorganic carbon fixation, fermentation, CH4 oxidation, and both starch/glycogen and peptidoglycan degradation decreased during succession, whereas several high efficiency processes, as well as CO oxidation and most types of OC degradation, increased.”
Chunxiang Hu, Professor, Institute of Hydrobiology, Chinese Academy of Sciences
Furthermore, utilizing co-occurrence networks, scientists found that the C-cycle in biocrusts consists of an assimilation module similar to primary production and a dissimilation module similar to secondary production. Researchers also discovered that during succession, dynamic changes in the connections between C-cycle pathways and microbial community composition took place.
“The two C-cycle modules were connected by the Calvin-Benson-Bassham cycle, ethanol and propionate fermentation, and they were balanced by drought and salinity,” Hu explained.
This research contributed to a better understanding of C-cycle pathways and regulatory frameworks in biocrust succession, as well as laying the foundations for potential multi-omics studies of these systems.
Source:
Journal reference:
Wang, Q., et al. (2022) Carbon cycle in the microbial ecosystems of biological soil crusts. Soil Biology and Biochemistry. doi.org/10.1016/j.soilbio.2022.108729.