People who have the same deletion of a section of chromosome 16 are susceptible to neurodevelopmental disorders; however, some may show moderate psychological symptoms like anxiety or depression, while others may have significant intellectual disability or developmental delay. How is this possible?
 Image credit: Andrii Yalanskyi/Shutterstock.com
Image credit: Andrii Yalanskyi/Shutterstock.com
To address this, a group of researchers from Penn State has created techniques to assess how genetic variations in other parts of a person's genome interact with the deletion to help predict the characteristics that person will exhibit. According to the researchers, this study shifts attention from the traditional focus on single causal variants leading to neurodevelopmental disorders to the role of interactions among a patient's entire genetic architecture and may help guide customized precision medical interventions for complex disorders.
The study was published in the journal Nature Communications.
“We know that many traits have complex genetic underpinnings, meaning that the way they appear is influenced by the interactions among many genes,” says Santhosh Girirajan, T. Ming Chu Professor of Genomics and head of the Department of Biochemistry and Molecular Biology in the Penn State Eberly College of Science, the leader of the research team.
We’ve been working for more than a decade with a deletion on chromosome 16, referred to as 16p12.1, that deletes eight genes and is associated with varying neurodevelopmental outcomes, including autism, developmental delay, and congenital abnormalities. We’ve studied the deletion in a fruit fly model and at the population level in humans, but we wanted to see if we could drill down to understand how clinical features of the deletion vary from family to family.
Santhosh Girirajan, Department of Biochemistry and Molecular Biology, Eberly College of Science, The Pennsylvania State University
According to the researchers, patients with the 16p12.1 deletion typically inherited it from a parent who carried the mutation but may not have had any clinical symptoms. This contrasts with many other genetic illnesses, which are frequently brought on by a new mutation in an individual.
A child that inherits the deletion from one of its parents gets half of its genome from the other parent. This new combination of genetic information in the child might expose the deletion to a different set of genetic variants – from the parent without the deletion – that it can interact with to cause different features. It also allows us to compare family members to help identify the secondary genetic variants that might influence the features that appear and their severity.
Santhosh Girirajan, Department of Biochemistry and Molecular Biology, Eberly College of Science, The Pennsylvania State University
The researchers employed two methods in an attempt to determine the reasons behind the differences between members of various families. Induced pluripotent stem cells (iPSC) from patients with the 16p12.1 deletion, their relatives, and healthy donors were employed in the first approach. The National Institute of Neurological Disorders and Stroke provided controls from healthy donors, and the patients' and their families' blood samples were used to create the iPSCs.
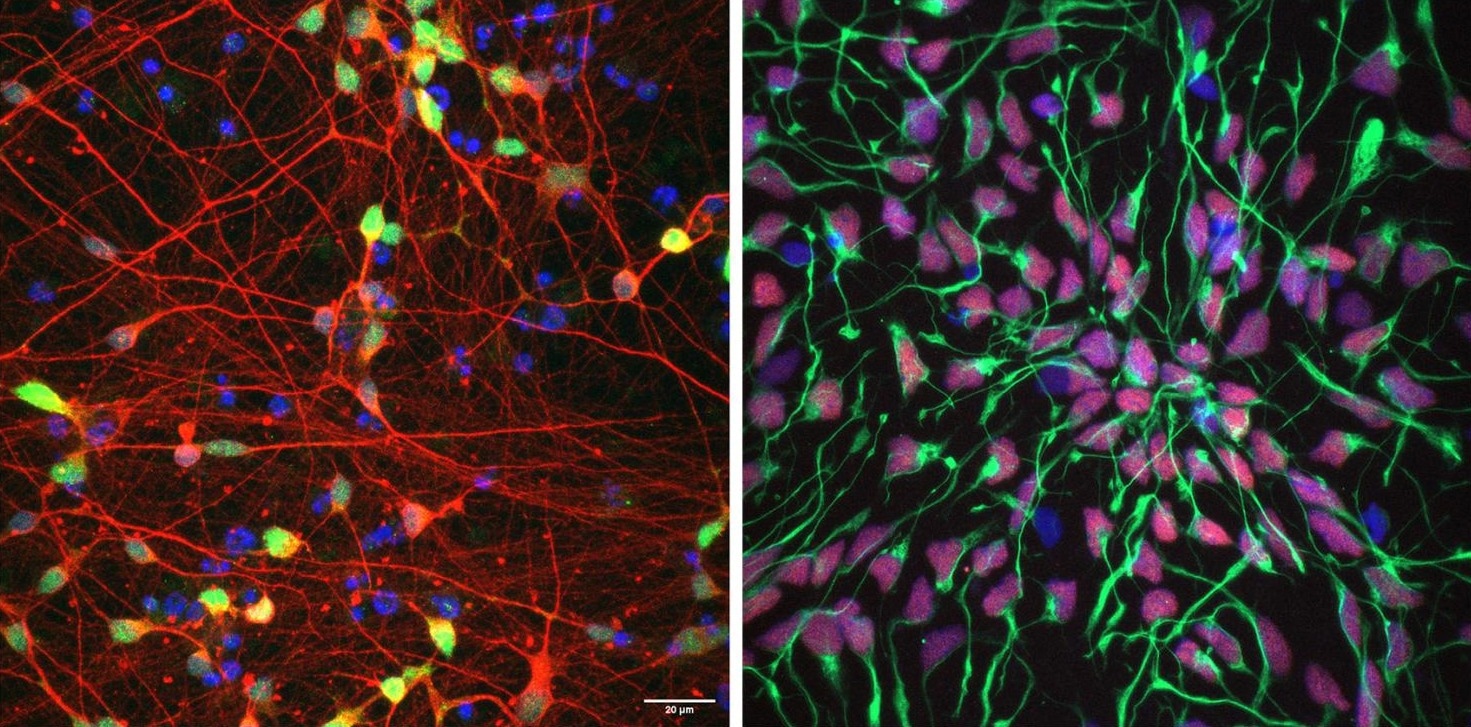 Researchers have developed a new framework to study the functional impact of genetic background on the expression and severity of neurodevelopmental features associated with a deletion of a portion of chromosome 16. The research team used induced pluripotent stem cells (iPSC) derived from patients with the deletion, their families and a healthy donor that they differentiated into different types of neuronal cells – image shows immunostained iPSC-derived neurons on the left and iPSC-derived neural progenitor cells on the right – to study how interactions among a patient’s entire genetic architecture and the deletion help determine the features that the individual will manifest. Image Credit: Girirajan Laboratory / Penn State.
Researchers have developed a new framework to study the functional impact of genetic background on the expression and severity of neurodevelopmental features associated with a deletion of a portion of chromosome 16. The research team used induced pluripotent stem cells (iPSC) derived from patients with the deletion, their families and a healthy donor that they differentiated into different types of neuronal cells – image shows immunostained iPSC-derived neurons on the left and iPSC-derived neural progenitor cells on the right – to study how interactions among a patient’s entire genetic architecture and the deletion help determine the features that the individual will manifest. Image Credit: Girirajan Laboratory / Penn State.
“Induced pluripotent stem cells (iPSCs), generated by reprogramming blood or skin cells, can be differentiated into many cell types. By adding certain small molecules, we can make them differentiate into different cell types in the neuronal lineage and then compare gene expression in cells from different families and cells with or without the deletion,” added Jiawan Sun.
The group transformed the iPSCs into mature and immature neurons as well as neural progenitor cells, which are precursors of neurons. Additionally, they induced the loss into iPSCs from a healthy donor using the CRISPR gene-editing technique. Individual differences in aberrant cell proliferation, cell death, and early differentiation were seen in several of these cell lines with the deletion.
According to the researchers, these anomalies are consistent with several clinical characteristics of people with the 16p12.1 deletion, such as differences in head size, which have already been documented in research on schizophrenia and autism.
The researchers discovered uncommon mutations in each person's genetic history by fully sequencing the genomes of every cell line utilized in the investigation. Additionally, they measured how much each gene was expressed in each type of cell. Based on this data, the team discovered that distinct variations in gene expression among the various cell lines were influenced by the genetic background of the iPSC lines. Additionally, they discovered variations in the cell lines' accessibility to genome regions that might regulate the expression of other genes but did not code for a gene.
There have been studies that compare cell lines with a particular mutation to otherwise identical cells without the mutation, and studies that look across a group of individuals looking for common genetic features shared by the group. What makes our study unique is the ability to identify variation in the genetic background that are family specific.
Santhosh Girirajan, Department of Biochemistry and Molecular Biology, Eberly College of Science, The Pennsylvania State University
In the cells from the healthy donor with the CRISPR-induced 16p12.1 deletion, the team also observed interactions with the genetic background. According to the researchers, this suggests that even otherwise healthy people may have genetic variations that, when combined with the deletion, may increase the chance of neurodevelopmental problems in their offspring.
Using CRISPR gene editing to gradually restore the function of each gene in the deletion was the team's second method for identifying variations in clinical characteristics linked to the 16p12.1 deletion. They discovered that every gene that was restored had an effect on the expression of a separate group of genes that varied throughout the study's cell types and families.
“We used to think the variation we see among individuals was caused by what we called the ‘two-hit’ model,” said Serena Noss. “The deletion was the first hit, and it could interact with a second-hit mutation elsewhere in the genome, but we’ve moved away from that language because we realize the deletion is interacting with multiple variants across an individual’s genome. It’s really more of a multi-hit model.”
According to the researchers, comprehending this multi-hit model and how a person's unique genetic architecture contributes to complicated neurodevelopmental disorders can aid in the creation of personalized treatments for these conditions.
Source:
Journal reference:
Sun, J., et al. (2026). Functional impact of genetic background on variable expressivity in neurodevelopmental disorders. Nature Communications. DOI: 10.1038/s41467-026-72598-z. https://www.nature.com/articles/s41467-026-72598-z.