Viruses are masters at entering human cells, thanks to specialized proteins that coat their surfaces. These viral surface proteins are frequently modified by scientists during vaccine development to investigate potential immune responses.
 Image credit: Spectral-Design/Shutterstock.com
Image credit: Spectral-Design/Shutterstock.com
However, synthetic proteins rarely behave as they would in a real virus, since they usually lack essential components of the virus's membrane. As a result, it has become challenging to understand how antibodies recognize and neutralize specific viral targets.
Researchers at Scripps Research have now developed a platform that enables the study of viral surface proteins in a manner that more closely mimics their natural appearance.
The novel method uses nanodisc technology, which preserves these proteins in a membrane-like state by embedding them within lipid particles. By better indicating the interaction between viral proteins and antibodies, this could aid in directing vaccine research.
According to a study published in Nature Communications, the platform was tested with proteins from HIV and Ebola, two viruses that have long posed challenges for vaccine researchers due to the immune system's inability to efficiently target their surface proteins. However, the method could be extended to other viruses such as SARS-CoV-2 and influenza that have similar membrane-embedded proteins.
For many years, we’ve had to rely on versions of viral proteins that are missing important pieces. Our platform lets us study these proteins in a setting that better reflects their natural environment, which is critical if we want to understand how protective antibodies recognize a virus.
William Schief, Co-Senior Author and Professor, Scripps Research
Surface proteins in real viruses are organized in particular shapes and embedded in a lipid membrane rather than floating freely. However, to facilitate the production and analysis of proteins, most laboratory investigations eliminate the membrane-anchoring region. Although helpful, this may mask essential information, especially for antibodies targeting regions near the viral membrane and the protein's base.
In this research, the team assembled vaccine candidate viral proteins into nanodiscs, small, stable membrane patches that hold the proteins in place. These lipid discs replicate the virus’s outer membrane, maintaining how antibodies recognize its proteins. The platform enabled researchers to use standard vaccine-development tools such as antibody-binding assays, immune cell sorting, and high-resolution imaging.
“Putting all of these components together into a single, reliable system was the key,” notes first author Kimmo Rantalainen, a senior scientist in Schief’s lab. “The individual pieces already existed, but making them work together in a way that’s reproducible and scalable opens up new possibilities for how vaccines are analyzed and designed.”
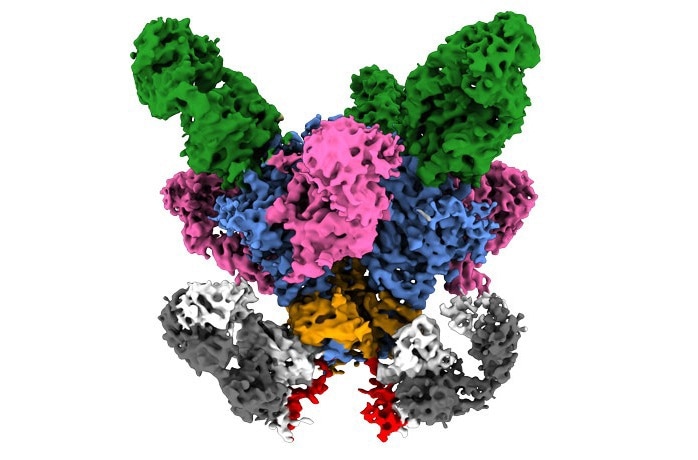 A viral surface protein (blue and orange) is shown bound to multiple antibodies (pink, green and gray/white), with a region near the membrane (red). Image Credit: Scripps Research
A viral surface protein (blue and orange) is shown bound to multiple antibodies (pink, green and gray/white), with a region near the membrane (red). Image Credit: Scripps Research
The scientists focused on a conserved region of the virus's surface protein close to the membrane, using HIV as a test case. A family of antibodies that can stop almost all HIV variations targets this area. These antibodies identify viral parts that remain identical even as they mutate – an immune response researchers hope vaccines might eventually stimulate.
Using their nanodisc platform, the researchers recorded in-depth structural snapshots of how these antibodies interact with the viral protein in its membrane context, revealing characteristics that are not apparent when the protein is examined separately. These discoveries also shed light on how some antibodies might kill a virus by disrupting the protein structures it uses to infect cells, providing hints on how future vaccines might more effectively elicit similar immune responses.
The structure gave us a level of detail we simply couldn’t access before. It showed us new interactions at the membrane interface and suggested why those matter for antibody function.
Kimmo Rantalainen, Scripps Research
The team also used their nanodisc platform to identify and bind Ebola proteins in the same membrane-like environment, demonstrating that the strategy is not limited to HIV.
This platform can also be used to investigate immunological responses to potential vaccines. Researchers can better understand how the body responds to a particular vaccine candidate by isolating and examining cells that recognize viral proteins, using nanodiscs as molecular "bait."
Additionally, because the system is scalable, preparations that formerly took a month or more may now be completed in roughly a week, making it feasible to compare several candidate designs side by side.
Scientists can use the platform to educate and expedite vaccine research, especially for viruses for which conventional methods have failed, even though it is not a vaccine in and of itself.
This gives the field a more realistic, accurate way to test ideas early on. By improving how we study viral proteins and antibody responses, we hope this platform will help advance next-generation vaccines against some of the world’s most challenging viruses.
William Schief, Co-Senior Author and Professor, Scripps Research
Source:
Journal reference:
Rantalainen, K., et al. (2026). Virus glycoprotein nanodisc platform for vaccine analytics. Nature Communications. DOI: 10.1038/s41467-026-68985-1. https://www.nature.com/articles/s41467-026-68985-1.